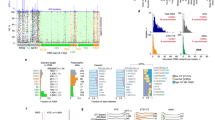
Abstract
The classical model of concerted evolution states that hundreds to thousands of ribosomal DNA (rDNA) units undergo homogenization, making the multiple copies of the individual units more uniform across the genome than would be expected given mutation frequencies and gene redundancy. While the universality of this over 50-year-old model has been confirmed in a range of organisms, advanced high throughput sequencing techniques have also revealed that rDNA homogenization in many organisms is partial and, in rare cases, even apparently failing. The potential underpinning processes leading to unexpected intragenomic variation have been discussed in a number of studies, but a comprehensive understanding remains to be determined. In this work, we summarize information on variation or polymorphisms in rDNAs across a wide range of taxa amongst animals, fungi, plants, and protists. We discuss the definition and description of concerted evolution and describe whether incomplete concerted evolution of rDNAs predominantly affects coding or non-coding regions of rDNA units and if it leads to the formation of pseudogenes or not. We also discuss the factors contributing to rDNA variation, such as interspecific hybridization, meiotic cycles, rDNA expression status, genome size, and the activity of effector genes involved in genetic recombination, epigenetic modifications, and DNA editing. Finally, we argue that a combination of approaches is needed to target genetic and epigenetic phenomena influencing incomplete concerted evolution, to give a comprehensive understanding of the evolution and functional consequences of intragenomic variation in rDNA.
Similar content being viewed by others
Introduction
Ribosomal DNAs (rDNAs) are best known for being the most conserved and heavily utilized house-keeping genes, encoding in eukaryotic organisms four types of structural ribosomal RNAs (rRNAs), i.e. 5S, 5.8S, 18S, and 28S/26S/25S rRNA. Three of these (5.8S, 18S, and 28S/26S/25S) are derived, respectively, from 18S-5.8S-28S/26S/25S rDNA (in this order) that are always linked in a single unit known as 45S rDNA in animals and 35S rDNA in plants. By convention, the nomenclature of rRNA/rDNA is derived from sedimentation rates of rRNA macromolecules which differ between the groups (Hemleben et al. 2021). The 45S/35S rDNAs will hereafter be referred as “45S rDNA(s)” and the 28S/26S/25S rDNAs as “28S rDNA(s)” for reading convenience. In eukaryotes, rDNAs form a multi-copy family of sequences organized in tandem repeats across one or several loci. The individual genes are separated by internal transcribed (ITS1 and ITS2) and intergenic (IGS) spacers (Fig. 1). The active 45S rRNA genes constitute important chromosomal landmarks called nucleolar organizer regions (NORs). In mammals and in most seed plants 5S rDNA, coding 5S rRNA, is separated from 45S rDNA at independent loci. However, a physical linkage of 5S with 45S genes is found in some plant groups (Garcia et al. 2009; Garcia and Kovařík 2013) especially in early diverging plants (Sousa et al. 2020; Wicke et al. 2011), some vertebrates (Davidian et al. 2022) and invertebrates (Drouin et al. 1987, 1992) and is commonly encountered amongst yeasts (Petes 1980; Szostak and Wu 1980).
The rDNA (green dots) at the nucleolar organizer regions (NOR) at interphase and on metaphase chromosomes represent rDNA loci, with each rDNA locus comprised of arrays of rDNA units (green squares) separated by intergenic spacer sequences (IGS). Each unit is comprised of the 18S, 5.8S, and 28S rDNA subunits, separated by the internally transcribed spacers (ITS). TSS, transcription start site.
In eukaryotic genomes, there is amazing sequence similarity between rDNA units observed within the genomes, indicating that each unit does not evolve independently from others, i.e. their evolution is concerted (Brown et al. 1972; Zimmer et al. 1980). The high similarity of units, i.e. homogenization, has been explained by repeated unequal crossovers (Dover 1982) and modeled in computer simulations almost 50 years ago, where mutations arising in repeats are erased over generations (Smith 1974). However, similarity does not mean complete sequence identity and in many natural situations we witness variable levels of unit-to-unit variation. Such variation has been reported in a broad range of eukaryotes, including plants (Chelomina et al. 2016; Harpke and Peterson 2006; Mayol and Rosselló 2001; Osuna-Mascaró et al. 2022; Shao et al. 2018; Vazquez 2019; Wang et al. 2016; Weitemier et al. 2015; Xiao et al. 2010), mammals (Robicheau et al. 2017), fish (Pinhal et al. 2011), insects (Keller et al. 2006) and fungi (Sultanov and Hochwagen 2022; West et al. 2014) (for a more complete list, see Supplementary Table 1). In some genomes, mutations in coding regions render rDNA units inactive. For example, humans carry c. 300 copies of 45S rDNA (1 C genome), out of which 14 (5%) are pseudogenes harboring various mutations (Robicheau et al. 2017) and the number of pseudogenized copies of 5S rDNA is even higher (Sørensen and Frederiksen 1991). In the plant Cycas revoluta, it has been estimated that the fraction of 45S rDNA pseudogenes is even larger, reaching almost five thousand copies, c. 50% of the total number of rDNA copies (Wang et al. 2016). Sequence variation between rDNA units has been widely used to determine phylogenetic inference (Poczai and Hyvonen 2010). However, rDNA polymorphisms can provide both invalid phylogenetic relationships (Won and Renner 2005) and overestimation of species diversity (Sun et al. 2013). On the other hand, they can be utilized to map evolutionary histories in hybrid species (Rauscher et al. 2004; West et al. 2014) and resolve cryptic introgressants (Garcia et al. 2020).
Extensive studies have investigated and speculated upon the underpinning causes of intragenomic rDNA variations in a range of organisms (reviewed, e.g. in Smirnov et al. 2021; Symonová 2019), but much information remains to be collated and further developed, in particular a need for complete sequencing through rDNA loci. In this work, we initially summarize studies showing intragenomic polymorphisms in rDNA in animal, fungi, and plant species, occurring widely across much eukaryotic diversity. We then discuss biological factors influencing concerted evolution and contributing to rDNA diversity. Finally, intra-array diversity, i.e. nucleotide variation between the units within a single array, is proposed as a hallmark of inefficient or even failing concerted evolution.
Intragenomic rDNA variations occur in divergent genera across the eukaryote tree-of-life
We collected publications by searching “intragenomic variation, non-concerted/incomplete concerted evolution of rDNA, rDNA polymorphism” in Google Scholar (up to 2022). This identified 136 records covering plants (32%), animals (36%), fungi (28%), protists (1%), and some prokaryotes (3%), and involving about 300 species in total (Supplementary Table 1). It is significant to note the following:
-
(i)
Frequent intragenomic variations of 45S rDNA were found in ITS subregions (57% of all reports) (Supplementary Table 2). This might explain the occurrence of poor support for branch positions when using ITS markers in phylogenetic studies (Poczai and Hyvonen 2010). For instance, Song et al. (2012) investigated 178 plant species and found that intragenomic variation of ITS2 was frequent, with an average of 35 variants in each species’ genome. Exceptionally high intragenomic polymorphism in ITS was reported in Mammillaria (Harpke and Peterson 2006), Asclepias (Weitemier et al. 2015), and Cycas (Xiao et al. 2010). The extent of ITS intragenomic diversity may vary significantly among genera and even species within the same genus (Vazquez 2019; Weitemier et al. 2015).
-
(ii)
Species with rDNA polymorphisms in plants, animals, and fungi (Supplementary Table 2) do not have close phylogenetic relationships, i.e., rDNA variations are scattered across eukaryote diversity and do not show apparent variation with genome size. Some genera had only a few variants (indicated as Single Nucleotide Polymorphisms, SNPs) in a few subregions, whereas other genera have extensive variation across the whole rDNA unit (Supplementary Table 1).
-
(iii)
Variation in rDNA coding regions was generally several-fold lower than that of non-coding regions of rDNA units (Stage and Eickbush 2007). This is explained by the fact that ITS (and IGS) is under relaxed selection compared to the coding regions, the latter with high functional constraints and lacking third codon redundancy as is found in protein coding genes. However, both 18S and 28S rRNA genes contain subdomains, termed “core” and “expansion” regions (Hancock and Dover 1988), differing in the degree of sequence uniformity. Variation in expansion regions is more frequent than in cores (Stage and Eickbush 2007) which exhibit purifying selection signatures (Sultanov and Hochwagen 2022). Of note, differences between coding and non-coding regions were more obvious when using high frequency variant call cut-offs rather than low-frequency call cut-offs, underlining the importance of variant calling parameters in data interpretation.
-
(iv)
The rDNA copy number can change rapidly over a few generations despite being similar, on average, between parents and offsprings (Rabanal et al. 2017 and reviwed in Kindelay and Maggert 2023; Salim and Gerton 2019). Indeed, shifts in copy number may significantly influence genome size such that it is visible to selection, as shown in the plant Arabidopsis thaliana, which has a relatively small genome (Long et al. 2013). In addition, the size of the array may influence non-ribosome-related functions of rDNA, such as the maintenance of genome integrity (Kobayashi 2008).
-
(v)
Finally, variation between units is not limited to 45S rDNA, but it also occurs at 5S rDNA loci (Kellogg and Appels 1995; Schneeberger and Cullis 1992; Stepanenko et al. 2022; Tynkevich et al. 2022). In humans and mice, the copy number of 5S and 45S rDNA (occupying separate loci) seems to be harmonized between populations/strains (Gibbons et al. 2015) while at the genus level both loci tend to undergo independent evolution (Fehrer et al. 2021; Mahelka et al. 2013; Volkov et al. 2017) though both are likely to be influenced by similar genetic processes, namely amplification, recombination and elimination.
Certainly, there is no doubt of the occurrence of intra and intergenomic variations in rDNA unit copies. However, many observations of “rDNA diversity” need to be interpreted with caution. For example, the employment of different technical approaches used to identify rDNA variation may lead to results that are differentially interpreted, and it is difficult to compare the conclusions of these studies. For example, it is unclear to what extent different methodologies used to quantify polymorphisms or variations in rDNAs impact the results, e.g., it is not easy to compare results from RFLP (restriction fragment length polymorphism), PCR-cloning, high throughput sequencing (short and long reads) and the different bioinformatic methods used (e.g. SNP calling parameters). Moreover, artifacts stemming from PCR and amplification processes induce mutations or chimeras and the extent of biased amplification of pseudogenes is not known (Cronn et al. 2002). High throughput sequencing is also not without its problems. For example, rDNA units are inherently GC-rich, and contain microsatellites and other tandem repeats, particularly in the IGS. All these features negatively influence sequencing efficiency, resulting in low coverage of rDNA and even erratic base calls (Fan et al. 2022; Guiblet et al. 2018). Thus, ideally, a common methodology needs to be employed to accurately assess and compare the efficiency of concerted evolution in individual species.
Factors leading to incomplete concerted evolution of rDNAs
The incomplete concerted evolution of multigenic families results from inefficient homogenization processes (Eickbush and Eickbush 2007; Liao 1999). Here we discuss possible biological processes (Fig. 2) that influence homogenization within rDNA.
Interspecific hybridization and whole genome duplication
Interspecific hybridization linked with subsequent whole genome duplication (WGD), termed allopolyploidy, immediately results in rDNA heterogeneity since parental genomes contain different rDNA variants. Persistence of parental variation generates intragenomic heterogeneity and “incomplete concerted evolution” phenotypes. Evidence for this comes from a range of studies. For example, in Saccharomyces cerevisiae, hybrid strains have mosaic-like genomes and, on average, a nearly 3-fold higher rDNA variation than non-hybrid strains with “clean” structured genomes (James et al. 2009). In the plant Malus toringoides, a hybrid descendent of M. transitoria and M. kansuensis, ITS sequences of both parents are retained (Liang et al. 2015). We should bear in mind that the timing and extent of genomic changes following hybridization and WGD events vary between species and that these can affect the extent of rDNA homogenization. Some genomic alternations take place immediately with the onset of genome merger or WGD, whereas others take many generations (Adams and Wendel 2005). For example, only two generations were needed to homogenize rDNA in Armeria hybrids (Fuertes Aguilar et al. 1999). Also, significant differences exist between allopolyploid populations of independent origin (Borowska-Zuchowska and Hasterok 2017; Kovarik et al. 2005; Lowe and Abbott 1996; Sochorova et al. 2016) and even between individuals of the same origin (Bao et al. 2010) suggesting that changes at rDNA loci may be astonishingly fast upon ‘genomic shock’ induced by interspecific hybridization. However, intragenomic variation in IGS seems to be never entirely removed, possibly because of the inherent instability of the elements in its sequence structure (Lunerova et al. 2017). The adaptive significance, if any, of variation in IGS is debatable, since except for promoters and splicing site regions, the function of IGS is mostly unknown or absent (Fedoroff 1979).
WGD generates a large rRNA gene dosage change in a newly formed polyploid. It can be hypothesized that rDNA loci are particularly sensitive to WGD since the number of active genes needs to be harmonized with cellular requirements and organism physiology. In support of this hypothesis, the physical elimination of rDNA loci (both 5S and 45S) following WGD is commonly encountered and well-documented in multiple animals (Gromicho et al. 2006; Knytl et al. 2023; Roco et al. 2021; Symonová et al. 2017a; Tagliavini et al. 1999; Ye et al. 2017) and plant (Garcia et al. 2017; Kotseruba et al. 2003; Lim et al. 2007; Volkov et al. 2007) polyploid systems. Significantly, in some cases, uniparental elimination of loci has occurred even in synthetic polyploid lineages (Guo and Han 2014; He et al. 2012; Malinska et al. 2010; Pontes et al. 2004). Such locus loss is actually reducing intragenomic rDNA heterogeneity following allopolyploidy and could potentially have adaptive significance.
Figure 3 outlines short (immediate) and later divergence events apparent in rDNA loci of hybrids and allopolyploids. Central early players in rDNA divergence are likely to be epigenetic mechanisms, which appear to have dual, contrasting roles. The epigenetic marks signposting active rDNA units (Fig. 3, right) with transcribed genes may contribute to array homogeneity by facilitating recombination and gene conversion. Active decondensed chromatin may also be vulnerable to DNA breaks leading to locus loss and rearrangements. In contrast, repressive epigenetic marks, such as methylcytosine (5 mC) and histone H3K9 methylation (Fig. 3, left) may not only stabilize rDNA silencing (NOR inactivity, termed nucleolar dominance in allopolyploids, reviewed in Borowska-Zuchowska et al. (2023) but may also inhibit recombination (Melamed-Bessudo and Levy 2012; Underwood et al. 2018) and sequence homogenization. Over time, 5 mC, either passively or actively, deaminates and converts into thymine (T) in what is thought to be a random process. A consequence is that recombination is further inhibited because of insufficient similarity between the units, manifesting in an incomplete concerted evolution phenotype.
Top: parental A and B arrays (thick horizontal arrows) are unified in a newly formed nucleus after a hybridization event. The arrays take either a non-concerted evolution route leading to decreased homogeneity and incomplete homogenization phenotypes (left) or a concerted evolution route leading to increased homogeneity and complete homogenization phenotypes (right). In the left panel, only array A is shown for simplicity. Heterochromatic marks such as dimethylation of histone H3 lysine 9 (H3K9m2) (ovals in left) and cytosine methylation (“m” in left) may contribute to increased array heterogeneity and its subsequent degeneration. The “T” in bold indicates C > T mutations.
Frequency of meiotic cycles and asexual reproduction
Organisms with asexual (apomictic) reproduction might be expected to show low frequencies of concerted evolution due to a reduced rate of meiotic crossovers (Pringle et al. 2000). In support of this hypothesis, some apomictic species display increased diversity of rDNA units compared to their sexual counterparts (Fehrer et al. 2009; Machackova et al. 2022), although in other systems, differences are not so pronounced (Zaveska Drabkova et al. 2009). Elsewhere, in parthenogenetic lizards (Hillis et al. 1991) and some plant species with prevalent vegetative propagation (e.g., strawberry, onion, iris) (Fredotovic et al. 2014; Hizume et al. 2002; Lim et al. 2007; Liu and Davis 2011) or impaired meiosis (e.g., dogroses, Herklotz et al. 2018) there are normal (high) levels of rDNA homogeneity. It should be mentioned that asexual and sexual modes of reproduction may occasionally be switched in plants and ‘a little bit of sex may help to avoid genomic decay and extinction of apomictic populations’ (Hojsgaard and Hörandl 2015). Thus, rDNA diversity in apomictic species can be explained by inherited variation from past hybridization events rather than having accumulated with apomixis (discussed further below). Nevertheless, in the crustacean genus Daphnia, different spectra of rDNA variants can arise within 90 generations of apomictic reproduction (McTaggart et al. 2007). These observations indicate that rDNA may also be a target of some form of somatic recombination. It has been hypothesized that the nucleolus serves as a niche for this process due to intensive transcription of 45S rDNA leading to double-strand breaks (Kovarik et al. 2008). The subsequent repair process may both increase or decrease array homogeneity (Sims et al. 2021).
The effects of life span, genetic drift and genome size
Life span may also play a role in homogenization frequency, since herbaceous plant lineages have ITS substitution rates almost twice as high as woody plants (Kay et al. 2006), which are much longer lived. There is also evidence that long-lived species such as gymnosperms display a high diversity of repeats in their genomes (Nystedt et al. 2013). Indeed, amongst gymnosperms, the cycads (genus Cycas) show extraordinary intragenomic rDNA heterogeneity, high rDNA copy numbers and high pseudogene content (Wang et al. 2016). It has been speculated that many gymnosperms have expanded genomes (>18 Gb/1 C on average) because of failing or reduced recombination processes that would otherwise remove non-functional DNA, especially retroelements (Leitch and Leitch 2012). Such a phenomenon might also maintain non-functional rDNA copies. However, some short-lived animals such as the grasshopper Podisma pedestris also show a high diversity of rDNA repeats in its large (18 Gb/1 C) genome (Keller et al. 2006). Species in Cycas and to a lesser extent Podisma pedestris, occur in relatively isolated populations today (both perhaps from much larger ancestral populations) where the effects of genetic drift are likely to be significant. These observations argue that long life spans, small population sizes, large genome sizes and infrequent meiotic cycles are associated with incomplete homogenization.
Developmental factors and premeiotic recombination
Although generally rDNA units show faithful Mendelian inheritance, newly amplified variants of IGS not present in parental lineages have been reported among siblings in animals (Cluster et al. 1987; Reeder et al. 1976) and in lineages of synthetic allopolyploid plants (Lin et al. 1985; Skalicka et al. 2003). While these case examples remain unexplained, studies in Xenopus laevis (African clawed frog) point to developmental effects. In this organism, primordial germ cells amplify huge amounts of extrachromosomal rDNA through recombination which has cell-to-cell sequence variation, especially in the IGS (Kalt and Gall 1974; Bird 1978). Although extrachromosomal rDNA copies are lost during development, some of these molecules can potentially recombine with chromosomal rDNA arrays at the premeiotic stage, giving rise to new variants (Haig 2021). Concerted evolution of these variants can then increase rDNA homogeneity within the cell and increase heterogeneity among them. Variable rDNA genotypes arising from such a process can then be subjected to cellular selection in development. Selection for preadult developmental variation in intergenic spacers of X chromosome-linked rDNA loci have also been proposed in Drosophila (Cluster et al. 1987) and certain rDNA unit length variants have been correlated with development rates. In this context, Haig (2021) proposed that non-coding IGS sequences are subject to positive intranuclear selection for persistence and spread through arrays. It will be interesting to investigate plants, which lack true germline and determine if somatic mutations are transmitted to the next generation.
The contribution of environmental factors to rDNA variation
There is some evidence that environmental factors may also contribute to the occurrence of rDNA variation, or perhaps rDNA variation may increase the environmental adaptability of certain organisms. This is because rDNA variation may influence rDNA transcription, which may in turn affect translation of protein coding genes and cell physiology (Kurylo et al. 2018). For instance, high level of intragenomic variation of rDNA sequences was found and postulated to be associated with adaptability to severe environments in extremotolerant and extremophilic microorganisms (Lopez-Lopez et al. 2007). Relationships between IGS variants and environmental factors such as drought, rainfall, soil composition and different habitats have also been documented in several plant systems (Cluster and Allard 1995; Govindaraju and Cullis 1992; Saghai-Maroof et al. 1984; Sharma et al. 2004) and reviewed in Nieto Feliner and Rosselló (2012). However, it remains to be determined if variation in non-coding regions of rDNA units modifies the transcriptional efficiency of units and if these have adaptive significance.
Genetic and epigenetic barriers interfering with rDNA homogenization processes
As discussed in the previous section, inhibition of recombination might lead to rDNA variation/polymorphism. Alternatively, the same outcome can be expected if the mutation rate overcomes the frequency of recombination. Indeed, individual units in tandem arrays are vulnerable to degeneration and loss of functionality, and can be blind to selection until they accumulate to such an extent that the fitness of the organism is impaired. We argue that concerted evolution functions as a correction mechanism, secondary to DNA repair, that has evolved to better control the fidelity of multigenic, tandemly repeated, families. This may be particularly significant in multicellular eukaryotic organisms whose genomes are overcrowded with repeats. It is likely that recombination would be most frequent in a homogeneous array than in an array that contains a diversity of repeats, since recombination is dependent on sequence identity. In other words, it can be hypothesized that the excision of deleterious variant(s) by recombination is more likely if they occur in a homogeneous array. This hypothesis is supported by several observations in plants, where species bearing a diversity of repeats in their genomes tend also to have tremendously expanded genome sizes (Novak et al. 2020; Nystedt et al. 2013). Based on this theoretical supposition, we suggest the following factors as potential barriers hindering homogenization and giving rise to subsequent incomplete concerted evolution (Fig. 4):
Chromosome barriers
The number of rDNA loci could impact rDNA homogenization, since there is evidence that interlocus homogenization is less frequent than intralocus homogenization. Consequently, species with fewer rDNA loci are more likely to display complete homogenization of units than those with more rDNA loci. Indeed, the relationship between intragenomic diversity and locus number has been demonstrated in a number of plant species including Asclepias (Weitemier et al. 2015), Arabidopsis (Copenhaver and Pikaard 1996; Riddle and Richards 2005), Nicotiana (Matyasek et al. 2012) and Ephedra (Wang et al. 2016). Nevertheless, fungi bearing relatively few rDNA copies in their genomes show considerable heterogeneity of rDNA (Supplementary Table 1), and incomplete homogenization may be driven by other processes in these species (discussed further below).
Homogenization of rDNA is also likely to be influenced by the chromosome location of rDNA loci (Brownell et al. 1983). In cotton and tobacco allotetraploids, terminally located 45S rDNA loci were homogenized to near completion by interlocus gene conversion (Lim et al. 2000; Volkov et al. 1999; Wendel et al. 1995). Of note, gene conversion seems to be the only mechanism reducing rDNA diversity without changing locus numbers. Either terminal positions could be (or are) more favorable to recombination, or other positions could be (or are) unfavorable to recombination, or both. Certainly, recombination rates are not equal across the genome. However, active sites of recombination differ between species. Members of the Mus genus (mouse) bear high number of 45S loci with homogenized rDNA that are almost exclusively at pericentromeric position of telocentric chromosomes (Cazaux et al. 2011). Also, in Northern pike (fish) the highly amplified and homogenized 5S rDNA occur at pericentromeric positions of most telocentric/acrocentric chromosomes (Symonová et al. 2017b). Potentially the tendency towards homogenization could be determined by the physical proximity of rDNA clusters to telomeres or centromeres. The position of the individual rDNA unit within the rDNA array may also influence their likelihood to be homogenized. The edge-located copies in a single rDNA cluster apparently do not undergo homogenization or undergo it at a lower frequency, resulting in their pseudogenization in humans (Robicheau et al. 2017) and wheat (Tulpova et al. 2022). Horizontal transfers of DNA may also introduce rDNA variation. In grasses, an rDNA horizontal transfer resulted in the fast erosion of an rDNA array and its colonization by transposable elements (Mahelka et al. 2017). Conversely, another recently described rDNA horizontal transfer from the eudicot Potentilla to the monocot Erythronium dens-canis generated an intact and even partially expressed rDNA array from Potentilla (Bartha et al. 2022).
Theoretical models predict that the time needed for array homogenization increases roughly linearly with the initial size of the array (Smith 1974). The number of rDNA copies within eukaryotic genomes is variable but is positively correlated with genome size (Prokopowich et al. 2003). Recombination, including unequal crossing-over, can result in one recombinant chromosome having more rDNA copies and the other fewer copies. Chromosomes that have too few rDNA units might be selected against, perhaps as a result of accumulated deleterious mutations or insufficient copy numbers of functional units. Indeed, in allohexaploid wheat bearing multiple rDNA loci, the very small A-genome loci are more heterogeneous compared to larger B and D genome loci (Tulpova et al. 2022). However, too large an rDNA array may be prone to deletion mediated by intralocus recombination between distal units. Perhaps, there might be an optimal size for the array functionality, given the frequency of and position of recombination in the species.
Processes related to homology searches in DNA repair/recombination also need to be considered. In the filamentous fungus Neurosporra crassa, 5S rDNA unit copies can be targeted by Repeat Induced Point mutations (RIP) in the premeiotic phase of the life cycle (Selker and Stevens 1985). Loci exposed to RIP exhibit high rates of G → A and C → T transitions, elevating sequence divergence. The mechanism of repeat recognition for RIP involves direct interactions between homologous double-stranded DNA (dsDNA) segments in somatic cells (Gladyshev and Kleckner 2017). It should be noted that, in contrast to animals and plants, 5S rDNA does not form typical tandems in N. crassa and units are dispersed across multiple chromosomes. Nevertheless, RIP may potentially account for an unusually high level (18–83%) of 5S rDNA pseudogenes in these filamentous fungi (Rooney and Ward 2005), giving rise to deleterious mutations that are reversed by new copies that appear, consistent with the birth-and-death model of multigene family evolution (Nei and Rooney 2005). Whether RIP (or analogous mechanisms) accounts for rDNA pseudogenization in other multicellular eukaryotes remains to be determined.
Collectively, physical barriers including the position and site numbers of rDNAs on the chromosomes, chromosome rearrangements, intrinsic structure of arrays and the number of rDNA copies within one locus, are more likely functioning together rather than independently, and all are likely to influence recombination and homogenization of rDNAs.
rDNA structural barriers
Natural sequence variation is found in each of the rDNA unit subregions in most species (Fig. 1), although it is typically several-fold higher in IGS than in the rest of the unit (Ambrose and Crease 2011; Draisma et al. 2012; Krawczyk et al. 2017; Lunerova et al. 2017). IGS regions can be very large (up to tens of kb in some species) and provide a natural niche for alien sequence insertions, e.g. tandem repeat (sub-repeat) and even functional 5S rRNA genes (Drouin et al. 1992; Galián et al. 2012). The GC-rich minisatellites residing in many species’ IGS can be particularly polymorphic, indeed in Cucurbita moschata (pumpkin) IGS displays both high levels of intra- and inter-array heterogeneity (Matyasek et al. 2019). Similarly, most IGS polymorphisms in human rDNA are located at CT and TG repeated sequences (Fan et al. 2022). Indeed, much intra-genomic variation in rDNA is primarily driven by structural elements residing in the IGS, especially involving short tandem repeats.
Epigenetic modifications of chromatin, such as DNA methylation and histone modifications might occur in a step-wise manner and influence for example DNA condensation, chromatin structure, ultimately affecting recombination and homogenization processes (Potapova and Gerton 2019). The differential condensation of active vs inactive rDNA chromatin, which is driven by epigenetic status of the rDNA units, will also impact rates of recombination and hence frequency of rDNA homogenization.
Effector gene barriers
A large number of genes control genetic recombination, transcription, epigenetic modification, and DNA repair. The involvement of these genes should be considered to better understand mechanisms of incomplete concerted evolution of rDNAs.
Recombination and transcription-relevant genes are critical for our understanding of concerted evolution given their role in unequal crossovers and/or gene conversion and in the transcriptional activity of NORs (Cockrell and Gerton 2022). In budding yeast, it is well known that mutations in the SIR2 (encoding histone deacetylase) and FOB1 (a replication fork blocking factor) genes respectively increase and decrease recombination within rDNA repeats (Gottlieb and Esposito 1989; Kobayashi and Horiuchi 1996). Both genes exhibit multiple functions in a cell and particularly SIR2 has been relatively well described. Briefly, SIR2 encodes an NAD + -dependent histone deacetylase that catalyzes and accelerates the de-acetylation of histones H3 and H4 (Blander and Guarente 2004). Hypoacetylation of histones, limiting the accessibility of chromatin, is a heterochromatin hallmark in a wide range of organisms, from yeast to humans. With hypoacetylation of histones, rDNA transcription is silenced and the sister chromatid recombination on rDNA sites is inhibited (Smith and Boeke 1997). This may prolong the persistence of silenced parental rDNAs after the interspecific hybridization (Kovarik et al. 2008). Finally, a relationship between DNA damage and rDNA instability is evidenced by experiments in Drosophila, where experimentally induced DNA breaks by I-CreI endonuclease altered the rDNA copy number (Paredes and Maggert 2009).
Cytosine deamination processes are an abundant source of genetic variability in eukaryotic cells (Duncan and Miller 1980). In particular, methylated cytosines residues are mutation hot spots since deamination of 5mC leads directly to T (C → T substitution) while deamination of C leads to C → U substitutions (Fig. 3). Thus, mutation load might be higher in densely methylated genomes than in genomes with no or low levels of methylation. Indeed, the C → T transitions are the most abundant SNPs seen in plants (Buckler et al. 1997) and grasshoppers (Keller et al. 2006) rDNA. Both plants (Meyer 2011) and grasshoppers (Robinson et al. 2011) also bear high level of methylation in their DNA.
Such unit divergence is likely to inhibit homology searches and recombination (Fig. 3, left). In animals, cellular cytosine deaminase also known as (Aid)/apolipoprotein B mRNA-editing enzyme (APOBEC3) family (Pecori et al. 2022) converts C to T (or 5 mC to T) giving rise to T/G mismatches. These mismatches can be recognized and ultimately removed by the methyl-CpG-binding domain 4 (Mbd4) glycosylase or thymine DNA glycosylase (Tdg) (Kunal et al. 2008). Hypothetically, increased cytosine deaminase activity or base excision repair (BER) defects would elevate mutation rates in rDNA leading to incomplete homogenization. In Arabidopsis thaliana active demethylation is performed by the activity of DNA glycosylase, mainly referred to as DEM/ROS1 family glycosylases and the BER pathway (Ikeda and Kinoshita 2009; Zhu 2009). Briefly, DNA glycosylases (DME/ROS1 family) involved in the BER process first recognize and then directly remove various substrates, including T/G mismatches that in most cases are generated during deamination (Baute and Depicker 2008). However, the relationship between the APOBEC3 activities, (methyl)cytosine deamination and rDNA pseudogenization remain to be established. It should also be mentioned that strategies dealing with mismatches caused by cytosine deamination may differ between plants and animals (Law and Jacobsen 2010).
Taken together, we suggest that screening of the genes above and comparing across a range of species their sequences and activities with the levels of rDNA polymorphisms, would lead to a better understanding of the processes leading to incomplete concerted evolution of rDNAs.
Conclusions and perspectives
In the vast majority of eukaryotes, evidence of concerted evolution is observed in rDNAs arrays. However, numerous examples of rDNA sequence and copy number variations observed in a wide range of genera and in different subregions of rDNA units raise several questions, which include: can unbiased criteria discriminate between complete, incomplete, and even failing concerted evolution? To what extent (threshold) can intragenomic variation/polymorphisms in rDNA units be attributed to “incomplete” concerted evolution? How might we discriminate variants that could go to fixation (functional) from those that contribute to array heterogeneity and are non-functional? How strong is selection along the different sequence categories of rDNA units? What is the level of unit-to-unit variation within active and non-active arrays? What is the relationship between rDNA copy number and nucleotide variation?
In order to address these questions, it will be essential to improve methodical approaches used for scoring rDNA variation within and across the arrays establishing clearly defined, biologically relevant threshold values, e.g., specific number of variations, SNPs per DNA sequence length. MinION technology, generating >200 kb reads, appears to be a suitable method for long-scale analyses of rDNA arrangement and unit structure (McKinlay et al. 2021), as was applied to the latest human T2T genome assembly including rDNA assembly (Nurk et al. 2022). However, its per-base error rate is still high (5–15%) (Istace et al. 2017) preventing unambiguous base calling. Therefore, assembling the highly repetitive rDNAs at least covering several 18S-5.8S-28S-IGS units (each of these “units” has a size of c. 10–50 kb in general) with lower sequencing errors, requires with expensive and high coverage sequencing. The HiFi PacBio technology (PacBio Biosciences, USA) generating long reads seems to overcome the accuracy problem (the claimed accuracy of >99% is comparable to short reads or Sanger sequencing) and could become a method of choice for determining single nucleotide variation within rDNA arrays. In the course of manuscript revision a near complete reconstruction of Arabidopsis thaliana 5S rDNA clusters by employing the above-mentioned approach was reported (Rabanal et al. 2022). We believe that the phenomenon and potential mechanisms of incomplete concerted evolution of rDNA identified here may only represent the tip of the iceberg to fully understand the evolution and functional diversity of rDNA in the future.
Change history
02 August 2023
A Correction to this paper has been published: https://doi.org/10.1038/s41437-023-00644-3
References
Adams KL, Wendel JF (2005) Polyploidy and genome evolution in plants. Curr Opin Plant Biol 8(2):135–141
Ambrose CD, Crease TJ (2011) Evolution of the nuclear ribosomal DNA intergenic spacer in four species of the Daphnia pulex complex. BMC Genet 12:13
Bao Y, Wendel JF, Ge S (2010) Multiple patterns of rDNA evolution following polyploidy in Oryza. Mol Phylogenet Evol 55(1):136–142
Bartha L, Mandakova T, Kovarik A, Bulzu PA, Rodde N, Mahelka V et al. (2022) Intact ribosomal DNA arrays of Potentilla origin detected in Erythronium nucleus suggest recent eudicot-to-monocot horizontal transfer. N. Phytol 235(3):1246–1259
Baute J, Depicker A (2008) Base excision repair and its role in maintaining genome stability. Crit Rev Biochem Mol Biol 43(4):239–276
Bird AP (1978) A study of early events in ribosomal gene amplification. Cold Spring Harb Symp Quant Biol 42:1179–1183
Blander G, Guarente L (2004) The Sir2 family of protein deacetylases. Annu Rev Biochem 73:417–435
Borowska-Zuchowska N, Hasterok R (2017) Epigenetics of the preferential silencing of Brachypodium stacei-originated 35S rDNA loci in the allotetraploid grass Brachypodium hybridum. Sci Rep 7(1):5260
Borowska-Zuchowska N, Mykhailyk S, Robaszkiewicz E, Matysiak N, Mielanczyk L, Wojnicz R et al. (2023) Switch them off or not: selective rRNA gene repression in grasses. Trends Plant Sci 28:661–672
Brown DD, Wensink PC, Jordan E (1972) Comparison of ribosomal DNAs of Xenopus laevis and Xenopus mulleri: the evolution of tandem genes. J Mol Biol 63(1):57–73
Brownell E, Krystal M, Arnheim N (1983) Structure and evolution of human and African Ape rDNA Pseudogenes. Mol Biol Evol 1(1):29–37
Buckler ES, Ippolito A, Holtsford TP (1997) The evolution of ribosomal DNA: divergent paralogues and phylogenetic implications. Genetics 145(3):821
Cazaux B, Catalan J, Veyrunes F, Douzery EJP, Britton-Davidian J (2011) Are ribosomal DNA clusters rearrangement hotspots? A case study in the genus Mus (Rodentia, Muridae). BMC Evol Biol 11:124
Chelomina GN, Rozhkovan KV, Voronova AN, Burundukova OL, Muzarok TI, Zhuravlev YN (2016) Variation in the number of nucleoli and incomplete homogenization of 18S ribosomal DNA sequences in leaf cells of the cultivated Oriental ginseng (Panax ginseng Meyer). J Ginseng Res 40(2):176–184
Cluster PD, Allard RW (1995) Evolution of ribosomal DNA (rDNA) genetic structure in colonial Californian populations of Avena-Barbata. Genetics 139(2):941–954
Cluster PD, Marinkovic D, Allard RW, Ayala FJ (1987) Correlations between development rates, enzyme activities, ribosomal DNA spacer-length phenotypes, and adaptation in Drosophila melanogaster. Proc Natl Acad Sci USA 84(2):610–614
Cockrell AJ, Gerton JL (2022) Nucleolar organizer regions as transcription-based scaffolds of nucleolar structure and function. Results Probl Cell Differ 70:551–580
Copenhaver GP, Pikaard CS (1996) Two-dimensional RFLP analyses reveal megabase-sized clusters of rRNA gene variants in Arabidopsis thaliana, suggesting local spreading of variants as the mode for gene homogenization during concerted evolution. Plant J 9(2):273–282
Cronn R, Cedroni M, Haselkorn T, Grover C, Wendel JF (2002) PCR-mediated recombination in amplification products derived from polyploid cotton. Theor Appl Genet 104(2-3):482–489
Davidian AG, Dyomin AG, Galkina SA, Makarova NE, Dmitriev SE, Gaginskaya ER (2022) 45S rDNA repeats of turtles and crocodiles harbor a functional 5S rRNA gene specifically expressed in Oocytes. Mol Biol Evol 39(1):msab324
Dover GA (1982) Molecular drive: a cohesive mode of species evolution. Nature 299:111–117
Draisma SGA, Eurlings MCM, Lim PE (2012) High intra-individual sequence variation in the nuclear rDNA LSU-5S intergenic spacer in the Sargassaceae (Fucales, Phaeophyceae). J Appl Phycol 24(6):1373–1379
Drouin G, Hofman JD, Doolittle WF (1987) Unusual ribosomal RNA gene organization in copepods of the genus Calanus. J Mol Biol 196(4):943–946
Drouin G, Sevigny JM, Mclaren IA, Hofman JD, Doolittle WF (1992) Variable arrangement of 5S ribosomal genes within the ribosomal DNA repeats of Arthropods. Mol Biol Evol 9(5):826–835
Duncan BK, Miller JH (1980) Mutagenic deamination of cytosine residues in DNA. Nature 287(5782):560–561
Eickbush TH, Eickbush DG (2007) Finely orchestrated movements: evolution of the ribosomal RNA genes. Genetics 175(2):477–485
Fan WJ, Eklund E, Sherman RM, Liu H, Pitts S, Ford B et al. (2022) Widespread genetic heterogeneity of human ribosomal RNA genes. RNA 28(4):478–492
Fedoroff NV (1979) On spacers. Cell 16(4):697–710
Fehrer J, Krak K, Chrtek Jr J (2009) Intra-individual polymorphism in diploid and apomictic polyploid hawkweeds (Hieracium, Lactuceae, Asteraceae): disentangling phylogenetic signal, reticulation, and noise. BMC Evol Biol 9:239
Fehrer J, Slavikova R, Pastova L, Josefiova J, Mraz P, Chrtek J et al. (2021) Molecular evolution and organization of ribosomal DNA in the Hawkweed Tribe Hieraciinae (Cichorieae, Asteraceae). Front Plant Sci 12:647375
Fredotovic Z, Samanic I, Weiss-Schneeweiss H, Kamenjarin J, Jang TS, Puizina J (2014) Triparental origin of triploid onion, Allium x cornutum (Clementi ex Visiani, 1842), as evidenced by molecular, phylogenetic and cytogenetic analyses. BMC Plant Biol 14:14
Fuertes Aguilar J, Rosselló JA, Nieto Feliner G (1999) Nuclear ribosomal DNA (nrDNA) concerted evolution in natural and artificial hybrids of Armeria (Plumbaginaceae). Mol Ecol 8:1341–1346
Galián JA, Rosato M, Rosselló JA (2012) Early evolutionary colocalization of the nuclear ribosomal 5S and 45S gene families in seed plants: evidence from the living fossil gymnosperm Ginkgo biloba. Heredity 108(6):640–646
Garcia S, Kovařík A (2013) Dancing together and separate again: gymnosperms exhibit frequent changes of fundamental 5S and 35S rRNA gene (rDNA) organization. Heredity 111(1):23–33
Garcia S, Kovarik A, Leitch AR, Garnatje T (2017) Cytogenetic features of rRNA genes across land plants: analysis of the Plant rDNA database. Plant J 89(5):1020–1030
Garcia S, Lim KY, Chester M, Garnatje T, Pellicer J, Valles J et al. (2009) Linkage of 35S and 5S rRNA genes in Artemisia (family Asteraceae): first evidence from angiosperms. Chromosoma 118(1):85–97
Garcia S, Borowska-Zuchowska N, Wendel JF, Ainouche M, Kuderova A, Kovarik A (2020) The utility of graph clustering of 5S ribosomal DNA homoeologs in plant allopolyploids, homoploid hybrids and cryptic introgressants. Front Plant Sci 11:41
Gibbons JG, Branco AT, Godinho SA, Yu S, Lemos B (2015) Concerted copy number variation balances ribosomal DNA dosage in human and mouse genomes. Proc Natl Acad Sci USA 112(8):2485–2490
Gladyshev E, Kleckner N (2017) DNA sequence homology induces cytosine-to-thymine mutation by a heterochromatin-related pathway in Neurospora. Nat Genet 49(6):887–894
Gottlieb S, Esposito RE (1989) A new role for a yeast transcriptional silencer gene, SIR2, in regulation of recombination in ribosomal DNA. Cell 56(5):771–776
Govindaraju DR, Cullis CA (1992) Ribosomal DNA variation among populations of a Pinus rigida Mill (pitch pine) ecosystem: I. Distribution of copy numbers. Heredity 69:133–140
Gromicho M, Coutanceau JP, Ozouf-Costaz C, Collares-Pereira MJ (2006) Contrast between extensive variation of 28S rDNA and stability of 5S rDNA and telomeric repeats in the diploid-polyploid Squalius alburnoides complex and in its maternal ancestor Squalius pyrenaicus (Teleostei, Cyprinidae). Chromosome Res 14(3):297–306
Guiblet WM, Cremona MA, Cechova M, Harris RS, Kejnovska I, Kejnovsky E et al. (2018) Long-read sequencing technology indicates genome-wide effects of non-B DNA on polymerization speed and error rate. Genome Res 28(12):1767–1778
Guo X, Han FP (2014) Asymmetric epigenetic modification and elimination of rDNA sequences by polyploidization in wheat. Plant Cell 26(11):4311–4327
Haig D (2021) Concerted evolution of ribosomal DNA: Somatic peace amid germinal strife: Intranuclear and cellular selection maintain the quality of rRNA. Bioessays 43(12):e2100179
Hancock JM, Dover GA (1988) Molecular coevolution among cryptically simple expansion segments of eukaryotic 26S/28S rRNAs. Mol Biol Evol 5(4):377–391
Harpke D, Peterson A (2006) Non-concerted ITS evolution in Mammillaria (Cactaceae). Mol Phylogenet Evol 41(3):579–593
He WG, Qin QB, Liu SJ, Li TL, Wang J, Xiao J et al. (2012) Organization and variation analysis of 5S rDNA in different ploidy-level hybrids of red crucian carp x topmouth culter. Plos One 7(6):e38976
Hemleben V, Grierson D, Borisjuk N, Volkov RA, Kovarik A (2021) Personal perspectives on plant ribosomal RNA genes research: From precursor-rRNA to molecular Evolution. Front Plant Sci 12:797348
Herklotz V, Kovarik A, Lunerova J, Lippitsch S, Groth M, Ritz CM (2018) The fate of ribosomal RNA genes in spontaneous polyploid dogrose hybrids [Rosa L. sect. Caninae (DC.) Ser.] exhibiting non-symmetrical meiosis. Plant J 94(1):77–90
Hillis DM, Moritz C, Porter CA, Baker RJ (1991) Evidence for biased gene conversion in concerted evolution of ribosomal DNA. Science 251(4991):308–310
Hizume M, Shibata F, Matsusaki Y, Garajova Z (2002) Chromosome identification and comparative karyotypic analyses of four Pinus species. Theor Appl Genet 105(4):491–497
Hojsgaard D, Hörandl E (2015) A little bit of sex matters for genome evolution in asexual plants. Front Plant Sci 6:82
Ikeda Y, Kinoshita T (2009) DNA demethylation: a lesson from the garden. Chromosoma 118(1):37–41
Istace B, Friedrich A, d’Agata L, Faye S, Payen E, Beluche O et al. (2017) de novo assembly and population genomic survey of natural yeast isolates with the Oxford Nanopore MinION sequencer. Gigascience 6(2):1–13
James SA, O’Kelly MJ, Carter DM, Davey RP, Van OA, Roberts IN (2009) Repetitive sequence variation and dynamics in the ribosomal DNA array of Saccharomyces cerevisiae as revealed by whole-genome resequencing. Genome Res 19(4):626–635
Kalt MR, Gall JG (1974) Observations on early germ cell development and premeiotic ribosomal DNA amplification in Xenopus laevis. J Cell Biol 62(2):460–472
Kay KM, Whittall JB, Hodges SA (2006) A survey of nuclear ribosomal internal transcribed spacer substitution rates across angiosperms: an approximate molecular clock with life history effects. BMC Evol Biol 6:36
Keller I, Chintauan-Marquier IC, Veltsos P, Nichols RA (2006) Ribosomal DNA in the grasshopper Podisma pedestris: escape from concerted evolution. Genetics 174(2):863–874
Kellogg EA, Appels R (1995) Intraspecific and interspecific variation in 5S RNA genes are decoupled in diploid wheat relatives. Genetics 140(1):325–343
Kindeley SM, Maggert KA (2023) Under the magnifying glass: The ups and downs of rDNA copy number. Semin Cell Dev Biol 136:13638–13648
Knytl M, Fornaini NR, Bergelov B, Gvozdik V, Cernohorska H, Kubickova S et al. (2023) Divergent subgenome evolution in the allotetraploid frog Xenopus calcaratus. Gene 851:146974
Kobayashi T (2008) A new role of the rDNA and nucleolus in the nucleus-rDNA instability maintains genome integrity. Bioessays 30(3):267–272
Kobayashi T, Horiuchi T (1996) A yeast gene product, Fob1 protein, required for both replication fork blocking and recombinational hotspot activities. Genes Cells 1(5):465–474
Kotseruba V, Gernand D, Meister A, Houben A (2003) Uniparental loss of ribosomal DNA in the allotetraploid grass Zingeria trichopoda (2n = 8). Genome 46:156–163
Kovarik A, Pires JC, Leitch AR, Lim KY, Sherwood AM, Matyasek R et al. (2005) Rapid concerted evolution of nuclear ribosomal DNA in two Tragopogon allopolyploids of recent and recurrent origin. Genetics 169(2):931–944
Kovarik A, Dadejova M, Lim YK, Chase MW, Clarkson JJ, Knapp S et al. (2008) Evolution of rDNA in Nicotiana allopolyploids: a potential link between rDNA homogenization and epigenetics. Ann Bot 101(6):815–823
Krawczyk K, Nobis M, Nowak A, Szczecinska M, Sawicki J (2017) Phylogenetic implications of nuclear rRNA IGS variation in Stipa L. (Poaceae). Sci Rep 7(1):11506
Kunal R, Huggins IJ, James SR, Karpf AR, Jones DA, Cairns BR (2008) DNA demethylation in zebrafish involves the coupling of a deaminase, a glycosylase, and gadd45. Cell 135(7):1201–1212
Kurylo CM, Parks MM, Juette MF, Zinshteyn B, Altman RB, Thibado JK et al. (2018) Endogenous rRNA sequence variation can regulate stress response gene expression and phenotype. Cell Rep 25(1):236–248
Law JA, Jacobsen SE (2010) Establishing, maintaining and modifying DNA methylation patterns in plants and animals. Nat Rev Genet 11(3):204–220
Leitch AR, Leitch IJ (2012) Ecological and genetic factors linked to contrasting genome dynamics in seed plants. N Phytol 194(3):629–646
Liang T, Tang JM, Si T, Ju L, Zhou ZQ (2015) ITS sequence variation and concerted evolution in the natural hybrid species Malus toringoides. Nord J Bot 33(1):109–119
Liao D (1999) Concerted evolution: molecular mechanism and biological implications. Am J Hum Genet 64(1):24–30
Lim KY, Matyasek R, Kovarik A, Leitch A (2007) Parental origin and genome evolution in the allopolyploid Iris versicolor. Ann Bot 100(2):219–224
Lim KY, Kovarik A, Matyasek R, Bezdek M, Lichtenstein CP, Leitch AR (2000) Gene conversion of ribosomal DNA in Nicotiana tabacum is associated with undermethylated, decondensed and probably active gene units. Chromosoma 109(3):161–172
Lin L-S, Ho TD, Harlan JR (1985) Rapid amplification and fixation of new restricition sites in the ribosomal DNA repeats in the derivatives of a cross between maize and Tripsacum dactyloides. Dev Genet 6:101–112
Liu B, Davis TM (2011) Conservation and loss of ribosomal RNA gene sites in diploid and polyploid Fragaria (Rosaceae). BMC Plant Biol 11:157
Long Q, Rabanal FA, Meng DZ, Huber CD, Farlow A, Platzer A et al. (2013) Massive genomic variation and strong selection in Arabidopsis thaliana lines from Sweden. Nat Genet 45(8):884–U218
Lopez-Lopez A, Benlloch S, Bonfa M, Rodriguez-Valera F, Mira A (2007) Intragenomic 16S rDNA divergence in Haloarcula marismortui is an adaptation to different temperatures. J Mol Evol 65(6):687–696
Lowe AJ, Abbott RJ (1996) Origins of the new allopolyploid species Senecio cambrensis (asteraceae) and its relationship to the Canary Islands endemic Senecio teneriffae. Am J Bot 83(10):1365–1372
Lunerova J, Renny-Byfield S, Matyasek R, Leitch A, Kovarik A (2017) Concerted evolution rapidly eliminates sequence variation in rDNA coding regions but not in intergenic spacers in Nicotiana tabacum allotetraploid. Plant Syst Evol 303(8):1043–1060
Machackova P, Majesky L, Hrones M, Bilkova L, Hribova E, Vasut RJ (2022) New insights into ribosomal DNA variation in apomictic and sexual Taraxacum (Asteraceae). Bot J Linn Soc 199(4):790–815
Mahelka V, Kopecky D, Baum BR (2013) Contrasting patterns of evolution of 45S and 5S rDNA families uncover new aspects in the genome constitution of the agronomically important grass Thinopyrum intermedium (Triticeae). Mol Biol Evol 30(9):2065–2086
Mahelka V, Krak K, Kopecky D, Fehrer J, Safar J, Bartos J et al. (2017) Multiple horizontal transfers of nuclear ribosomal genes between phylogenetically distinct grass lineages. Proc Natl Acad Sci USA 114(7):1726–1731
Malinska H, Tate JA, Matyasek R, Leitch AR, Soltis DE, Soltis PS et al. (2010) Similar patterns of rDNA evolution in synthetic and recently formed natural populations of Tragopogon (Asteraceae) allotetraploids. BMC Evol Biol 10:291
Matyasek R, Kuderova A, Kutilkova E, Kucera M, Kovarik A (2019) Intragenomic heterogeneity of intergenic ribosomal DNA spacers in Cucurbita moschata is determined by DNA minisatellites with variable potential to form non-canonical DNA conformations. DNA Res 26(3):273–286
Matyasek R, Renny-Byfield S, Fulnecek J, Macas J, Grandbastien MA, Nichols R et al. (2012) Next generation sequencing analysis reveals a relationship between rDNA unit diversity and locus number in Nicotiana diploids. BMC Genom 13:722
Mayol M, Rossello JA (2001) Why nuclear ribosomal DNA spacers (ITS) tell different stories in Quercus. Mol Phylogenet Evol 19(2):167–176
McKinlay A, Fultz D, Wang F, Pikaard CS (2021) Targeted enrichment of rRNA gene tandem arrays for ultra-long sequencing by selective restriction endonuclease digestion. Front Plant Sci 12:656049
McTaggart S, Dudycha JL, Omilian A, Crease TJ (2007) Rates of recombination in the ribosomal DNA of apomictically propagated Daphnia obtusa lines. Genetics 175(1):311–320
Melamed-Bessudo C, Levy AA (2012) Deficiency in DNA methylation increases meiotic crossover rates in euchromatic but not in heterochromatic regions in Arabidopsis. Proc Natl Acad Sci USA 109(16):E981–E988
Meyer P (2011) DNA methylation systems and targets in plants. Febs Lett 585(13):2008–2015
Nei M, Rooney AP (2005) Concerted and birth-and-death evolution of multigene families. Annu Rev Genet 39:121–152
Nieto Feliner G, Rossello JA (2012) Concerted evolution of multigene families and homeologous recombination. In: Wendel JF (ed) Plant Genome Diversity(V1). Springer-Verlag: Wien. (pp:171–194).
Novak P, Guignard MS, Neumann P, Kelly LJ, Mlinarec J, Koblizkova A et al. (2020) Repeat-sequence turnover shifts fundamentally in species with large genomes. Nat Plants 6(11):1325–1329
Nurk S, Koren S, Rhie A, Rautiainen M, Bzikadze AV, Mikheenko A et al. (2022) The complete sequence of a human genome. Science 376(6588):44–53
Nystedt B, Street NR, Wetterbom A, Zuccolo A, Lin YC, Scofield DG et al. (2013) The Norway spruce genome sequence and conifer genome evolution. Nature 497(7451):579–584
Osuna-Mascaró C, de Casas RR, Berbel M, Gómez JM, Perfectti F (2022) Lack of ITS sequence homogenization in Erysimum species (Brassicaceae) with different ploidy levels. Sci Rep 2(1):16907
Paredes, KA, Maggert KA (2009) Expression of I-CreI endonuclease generates deletions within the rDNA of Drosophila Genetics, 181 (4): 1661–1671
Pecori R, Di Giorgio S, Lorenzo JP, Papavasiliou FN (2022) Functions and consequences of AID/APOBEC-mediated DNA and RNA deamination. Nat Rev Genet 23(8):505–518
Petes TD (1980) Unequal meiotic recombination within tandem arrays of yeast ribosomal DNA genes. Cell 19(3):765–774
Pinhal D, Yoshimura TS, Araki CS, Martins C (2011) The 5S rDNA family evolves through concerted and birth-and-death evolution in fish genomes: an example from freshwater stingrays. BMC Evol Biol 11:151
Poczai P, Hyvonen J (2010) Nuclear ribosomal spacer regions in plant phylogenetics: problems and prospects. Mol Biol Rep 37(4):1897–1912
Pontes O, Neves N, Silva M, Lewis MS, Madlung A, Comai L et al. (2004) Chromosomal locus rearrangements are a rapid response to formation of the allotetraploid Arabidopsis suecica genome. Proc Natl Acad Sci USA 101(52):18240–18245
Potapova TA, Gerton JL (2019) Ribosomal DNA and the nucleolus in the context of genome organization. Chromosome Res 27(1-2):109–127
Pringle A, Moncalvo JM, Vilgalys R (2000) High levels of variation in ribosomal DNA sequences within and among spores of a natural population of the arbuscular mycorrhizal fungus Acaulospora colossica. Mycologia 92(2):259–268
Prokopowich CD, Gregory TR, Crease TJ (2003) The correlation between rDNA copy number and genome size in eukaryotes. Genome 46(1):48–50
Rabanal FA, Nizhynska V, Mandakova T, Novikova PY, Lysak MA, Mott R et al. (2017) Unstable inheritance of 45S rRNA genes in Arabidopsis thaliana. G3 (Bethesda) 7(4):1201–1209
Rabanal FA, Gräff M, Lanz C, Fritschi K, Llaca V, Lang M et al. (2022) Pushing the limits of HiFi assemblies reveals centromere diversity between two Arabidopsis thaliana genomes. Nucleic Acids Res 50(21):12309–12327
Rauscher JT, Doyle JJ, Brown AH (2004) Multiple origins and nrDNA internal transcribed spacer homeologue evolution in the Glycine tomentella (Leguminosae) allopolyploid complex. Genetics 166(2):987–998
Reeder RH, Brown DD, Wellauer PK, Dawid IB (1976) Patterns of ribosomal DNA spacer lengths are inherited. J Mol Biol 105(4):507–516
Riddle NC, Richards EJ (2005) Genetic variation in epigenetic inheritance of ribosomal RNA gene methylation in Arabidopsis. Plant J 41(4):524–532
Robicheau BM, Susko E, Harrigan AM, Snyder M (2017) Ribosomal RNA genes contribute to the formation of pseudogenes and junk DNA in the human genome. Genome Biol Evol 9(2):380–397
Robinson KL, Tohidi-Esfahani D, Lo N, Simpson SJ, Sword GA (2011) Evidence for widespread genomic methylation in the migratory locust, Locusta migratoria (Orthoptera: Acrididae). Plos One 6(12):e28167
Roco AS, Liehr T, Ruiz-Garcia A, Guzman K, Bullejos M (2021) Comparative distribution of repetitive sequences in the karyotypes of Xenopus tropicalis and Xenopus laevis (Anura, Pipidae). Genes 12(5):617
Rooney AP, Ward TJ (2005) Evolution of a large ribosomal RNA multigene family in filamentous fungi: birth and death of a concerted evolution paradigm. Proc Natl Acad Sci USA 102(14):5084–5089
Saghai-Maroof MA, Soliman KM, Jorgensen RA, Allard RW (1984) Ribosomal DNA spacer-length polymorphisms in barley: mendelian inheritance, chromosomal location, and population dynamics. Proc Natl Acad Sci USA 81(24):8014–8018
Salim D, Gerton JL (2019) Ribosomal DNA instability and genome adaptability Chromosome Research 27(1–2):73–87
Schneeberger RG, Cullis CA (1992) Intraspecific 5S ribosomal RNA gene variation in flax, Linum-Usitatissimum (Linaceae). Plant Syst Evol 183(3-4):265–280
Selker EU, Stevens JN (1985) DNA methylation at asymmetric sites is associated with numerous transition mutations. Proc Natl Acad Sci USA 82(23):8114–8118
Shao Y, Zhang M, Xu Y, Zhu YQ, Yonezawa T, Wang YG et al. (2018) An improved metagenomic strategy reveals an unprecedentedly high level of intragenomic polymorphism of ribosomal DNA in three species of Camellia. J Syst Evol 56(3):250–258
Sharma S, Beharav A, Balyan HS, Nevo E, Gupta PK (2004) Ribosomal DNA polymorphism and its association with geographical and climatic variables in 27 wild barley populations from Jordan. Plant Sci 166(2):467–477
Sims J, Rabanal FA, Elgert C, von Haeseler A, Schlögelhofer P (2021) It is just a matter of time: balancing homologous recombination and non-homologous end joining at the rDNA locus during meiosis. Front Plant Sci 12:773052
Skalicka K, Lim KY, Matyasek R, Koukalova B, Leitch AR, Kovarik A (2003) Rapid evolution of parental rDNA in a synthetic tobacco allotetraploid line. Am J Bot 90(7):988–996
Smirnov E, Chmurciakova N, Liska F, Bazantova P, Cmarko D (2021) Variability of human rDNA. Cells 10(2):196
Smith GP (1974) Unequal crossover and the evolution of multigene families. Cold Spring Harb Symp Quant Biol 38:507–513
Smith JS, Boeke JD (1997) An unusual form of transcriptional silencing in yeast ribosomal DNA. Genes Dev 11(2):241–254
Sochorova J, Coriton O, Kuderova A, Lunerova J, Chevre AM, Kovarik A (2016) Gene conversion events and variable degree of homogenization of rDNA loci in cultivars of Brassica napus. Ann Bot 119:13–26
Song JY, Shi LC, Li DZ, Sun YZ, Niu YY, Chen ZD et al. (2012) Extensive pyrosequencing reveals frequent intra-genomic variations of internal transcribed spacer regions of nuclear ribosomal DNA. Plos One 7(8):e43971
Sørensen PD, Frederiksen S (1991) Characterization of human 5S rRNA genes. Nucleic Acids Res 19(15):4147–4151
Sousa A, Bechteler J, Temsch EM, Renner SS (2020) Different from tracheophytes, liverworts commonly have mixed 35S and 5S arrays. Ann Bot 125(7):1057–1064
Stage DE, Eickbush TH (2007) Sequence variation within the rRNA gene loci of 12 Drosophila species. Genome Res 17(12):1888–1897
Stepanenko A, Chen GM, Hoang PTN, Fuchs J, Schubert I, Borisjuk N (2022) The Ribosomal DNA loci of the ancient monocot Pistia stratiotes L. (Araceae) contain different variants of the 35S and 5S ribosomal RNA gene units. Front Plant Sci 13:819750
Sultanov D, Hochwagen A (2022) Varying strength of selection contributes to the intragenomic diversity of rRNA genes. Nat Commun 13(1):7245
Sun DL, Jiang X, Wu QLL, Zhou NY (2013) Intragenomic heterogeneity of 16S rRNA genes causes overestimation of prokaryotic diversity. Appl Environ Micro 79(19):5962–5969
Symonová R (2019) Integrative rDNAomics- Importance of the oldest repetitive fraction of the eukaryote genome. Genes 10(5):345
Symonová R, Havelka M, Amemiya CT, Howell WM, Korinkova T, Flajshans M et al. (2017a) Molecular cytogenetic differentiation of paralogs of Hox paralogs in duplicated and re-diploidized genome of the North American paddlefish (Polyodon spathula). BMC Genet 18(1):19
Symonová R, Ocalewicz K, Kirtiklis L, Delmastro GB, Pelikanova S, Garcia S et al. (2017b) Higher-order organisation of extremely amplified, potentially functional and massively methylated 5S rDNA in European pikes (Esox sp.). BMC Genomics 18(1):391
Szostak JW, Wu R (1980) Unequal crossing over in the ribosomal DNA of Saccharomyces cerevisiae. Nature 284(5755):426–430
Tagliavini J, Williot P, Congiu L, Chicca M, Lanfredi M, Rossi R et al. (1999) Molecular cytogenetic analysis of the karyotype of the European Atlantic sturgeon, Acipenser sturio. Heredity 83:520–525
Tulpova Z, Kovarik A, Toegelova H, Navratilova P, Kapustova V, Hribova E et al. (2022) Fine structure and transcription dynamics of bread wheat ribosomal DNA loci deciphered by a multi-omics approach. Plant Genome 15(1):e20191
Tynkevich YO, Shelyfist AY, Kozub LV, Hemleben V, Panchuk II, Volkov RA (2022) 5S ribosomal DNA of genus Solanum: molecular organization, evolution, and taxonomy. Front Plant Sci 13:852406
Underwood CJ, Choi K, Lambing C, Zhao XH, Serra H, Borges F et al. (2018) Epigenetic activation of meiotic recombination near Arabidopsis thaliana centromeres via loss of H3K9me2 and non-CG DNA methylation. Genome Res 28(4):519–531
Vazquez ML (2019) Molecular evolution of the internal transcribed spacers in red oaks (Quercus sect. Lobatae). Comput Biol Chem 83:107117
Volkov RA, Komarova NY, Hemleben V (2007) Ribosomal DNA in plant hybrids: inheritance, rearrangement, expression. Syst Biodivers 5:261–276
Volkov RA, Borisjuk NV, Panchuk II, Schweizer D, Hemleben V (1999) Elimination and rearrangement of parental rDNA in the allotetraploid Nicotiana tabacum. Mol Biol Evol 16(3):311–320
Volkov RA, Panchuk II, Borisjuk NV, Hosiawa-Baranska M, Maluszynska J, Hemleben V (2017) Evolutional dynamics of 45S and 5S ribosomal DNA in ancient allohexaploid Atropa belladonna. BMC Plant Biol 17(1):21
Wang W, Ma L, Becher H, Garcia S, Kovarikova A, Leitch IJ et al. (2016) Astonishing 35S rDNA diversity in the gymnosperm species Cycas revoluta Thunb. Chromosoma 125(4):683–699
Weitemier K, Straub SCK, Fishbein M, Liston A (2015) Intragenomic polymorphisms among high-copy loci: a genus-wide study of nuclear ribosomal DNA in Asclepias (Apocynaceae). Peer J 3:e718
Wendel JF, Schnabel A, Seelanan T (1995) Bidirectional interlocus concerted evolution following allopolyploid speciation in cotton (Gossypium). Proc Natl Acad Sci USA 92(1):280–284
West C, James SA, Davey RP, Dicks J, Roberts IN (2014) Ribosomal DNA sequence heterogeneity reflects intraspecies phylogenies and predicts genome structure in two contrasting yeast species. Syst Biol 63(4):543–554
Wicke S, Costa A, Munoz J, Quandt D (2011) Restless 5S: the re-arrangement(s) and evolution of the nuclear ribosomal DNA in land plants. Mol Phylogenet Evol 61(2):321–332
Won H, Renner SS (2005) The internal transcribed spacer of nuclear ribosomal DNA in the gymnosperm Gnetum. Mol Phylogenet Evol 36(3):581–597
Xiao LQ, Moller M, Zhu H (2010) High nrDNA ITS polymorphism in the ancient extant seed plant Cycas: incomplete concerted evolution and the origin of pseudogenes. Mol Phylogenet Evol 55(1):168–177
Ye LH, Zhang C, Tang XJ, Chen YY, Liu SJ (2017) Variations in 5S rDNAs in diploid and tetraploid offspring of red crucian carp x common carp. BMC Genet 18(1):75
Zaveska Drabkova L, Kirschner J, Stepanek J, Zavesky L, Vlcek C (2009) Analysis of nrDNA polymorphism in closely related diploid sexual, tetraploid sexual and polyploid agamospermous species. Plant Syst Evol 278(1-2):67–85
Zhu JK (2009) Active DNA demethylation mediated by DNA glycosylases. Annu Rev Genet 43(1):143
Zimmer EA, Martin SL, Beverley SM, Kan YW, Wilson AC (1980) Rapid duplication and loss of genes coding for the alpha chains of hemoglobin. Proc Natl Acad Sci USA 77(4):2158–2162
Acknowledgements
We thank the Xinglin Fund of Guangzhou University of Chinese Medicine (A1-AFD018191Z0145), Czech Science Foundation (22–16826S), Innovation Project for Forestry Science and Technology in Guangdong (2021KJCX015), as well as the Agencia Estatal de Investigación (PID2020-119163GB-I00 funded by MCIN/AEI/10.13039/501100011033) for support. We apologize to all authors whose works are cited in the Supplementary Tables and which could not be included in the main text from space reasons. Finally we wish to thank Frank Hailer, an evolutionary biologist from Cardiff University, UK, and three anonymous reviewers for their constructive comments.
Author information
Authors and Affiliations
Contributions
WW and AK conceived and wrote the manuscript, all the authors revised and proofed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Associate editor: Frank Hailer.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, W., Zhang, X., Garcia, S. et al. Intragenomic rDNA variation - the product of concerted evolution, mutation, or something in between?. Heredity 131, 179–188 (2023). https://doi.org/10.1038/s41437-023-00634-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41437-023-00634-5