Abstract
Purpose
Common scab of potatoes (CS) is influenced by plant-microbe-soil interactions, which are locally specific. The study aims to identify any common trends in such interactions across a landscape differentiated by soil pH and CS severity.
Methods
The 33 sites and 17 cultivars were characterized using soil and periderm nutrient contents and microbial communities. Quantitative PCR and Illumina amplicon sequencing were used to assess abundance of bacteria, actinobacteria and pathogens, and community composition.
Results
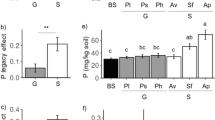
Comparisons between bulk and tuberosphere soil compartments as well as potato cultivars divided to three categories of CS susceptibility revealed that nitrogen was elevated in tuberosphere soil and N, Mg and Fe were lowered in periderm of resistant cultivars. The susceptible cultivar Agria grown at 7 sites had higher Ca content in tuberosphere soil, while the resistant cultivar Adela grown at 10 sites had higher S, P and Mg contents in its tuberosphere soil and P and Fe in periderm. That suggests further interactions between plants and bacterial community involving nutrient uptake. Diversity of bacteria was positively correlated with CS severity suggesting interactions between the Streptomyces pathogen populations and the local soil community.
Conclusions
Overall, pathogen abundance assessed by quantifying the thaxtomin biosynthetic txtB genes were randomly dispersed among the sites without connections to CS severity or soil pH. Thus, the significant differences between bacterial communities of bulk and tuberosphere soils together with cultivar CS susceptibility showed that the susceptible cultivars select bacterial community relatively similar to the bulk soil, while the resistant cultivars promote more distinct communities.





Similar content being viewed by others
Data Availability
Illumina MiSeq amplicon sequences of bacterial 16 S rRNA genes are available in the NCBI Sequence Read Archive (www.ncbi.nlm.nih.gov/sra) as BioProjects PRJNA699043 and PRJNA474544. All other data is available in the Supplementary Material.
References
Bogunovic I, Pereira P, Brevik EC (2017) Spatial distribution of soil chemical properties in an organic farm in Croatia. Sci Total Environ 584–585:535–545. https://doi.org/10.1016/j.scitotenv.2017.01.062
Braun S, Gevens A, Charkowski A et al (2017) Potato common scab: a review of the causal pathogens, management practices, varietal resistance screening methods, and host resistance. Am J Potato Res 94:283–296. https://doi.org/10.1007/s12230-017-9575-3
Caporaso JG, Lauber CL, Walters WA et al (2011) Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc Natl Acad Sci U S A 108(Suppl):4516–4522. https://doi.org/10.1073/pnas.1000080107
Cha JY, Han S, Hong HJ et al (2016) Microbial and biochemical basis of a Fusarium wilt-suppressive soil. ISME J 10:119–129. https://doi.org/10.1038/ismej.2015.95
Clarke CR, Kramer CG, Kotha RR et al (2019) Cultivar resistance to common scab disease of potato is dependent on the pathogen species. Phytopathology 109:1544–1554. https://doi.org/10.1094/PHYTO-09-18-0368-R
Curless MA, Kelling KA, Speth PE et al (2012) Effect of manure application timing on potato yield, quality, and disease incidence. Am J Potato Res 89:363–373. https://doi.org/10.1007/s12230-012-9256-1
Davis JR, McDole RE, Callihan RH (1976) Fertilizer effects on common scab of potato and the relation of calcium and phosphate-phosphorus. Phytopathology 66:1236–1241
Dees MW, Wanner LA (2012) In Search of Better Management of Potato Common Scab. Potato Res 55:249–268. https://doi.org/10.1007/s11540-012-9206-9
Dees MW, Sletten A, Hermansen A (2013) Isolation and characterization of Streptomyces species from potato common scab lesions in Norway. Plant Pathol 62:217–225. https://doi.org/10.1111/j.1365-3059.2012.02619.x
Edgar RC (2013) UPARSE: Highly accurate OTU sequences from microbial amplicon reads. Nat Methods 10:996–998. https://doi.org/10.1038/nmeth.2604
Fierer N (2017) Embracing the unknown: Disentangling the complexities of the soil microbiome. Nat Rev Microbiol 15:579–590. https://doi.org/10.1038/nrmicro.2017.87
Fierer N, Jackson RB (2006) The diversity and biogeography of soil bacterial communities. Proc Natl Acad Sci U S A 103:626–631. https://doi.org/10.1073/pnas.0507535103
Fiers M, Edel-Hermann V, Chatot C et al (2012) Potato soil-borne diseases. A review. Agron Sustain Dev 32:93–132. https://doi.org/10.1007/s13593-011-0035-z
Fofana B, Somalraju A, Fillmore S et al (2020) Comparative transcriptome expression analysis in susceptible and resistant potato (Solanum tuberosum) cultivars to common scab (Streptomyces scabies) revealed immune priming responses in the incompatible interaction. PLoS One 15:1–27. https://doi.org/10.1371/journal.pone.0235018
Glick BR (2012) Plant growth-promoting bacteria: mechanisms and applications. Scientifica (Cairo) 2012:1–15. https://doi.org/10.6064/2012/963401
Gu Y, Wei Z, Wang X et al (2016) Pathogen invasion indirectly changes the composition of soil microbiome via shifts in root exudation profile. Biol Fertil Soils 52:997–1005. https://doi.org/10.1007/s00374-016-1136-2
Hausvater E, Dolezal P (2013) Actinobacterial common scab of potatoes [in Czech]. Potato Research Institute, Havlíčkův Brod
Inceoǧlu Ö, Salles JF, van Elsas JD (2012) Soil and cultivar type shape the bacterial community in the potato rhizosphere. Microb Ecol 63:460–470. https://doi.org/10.1007/s00248-011-9930-8
Johnson EG, Joshi MV, Gibson DM, Loria R (2007) Cello-oligosaccharides released from host plants induce pathogenicity in scab-causing Streptomyces species. Physiol Mol Plant Pathol 71:18–25. https://doi.org/10.1016/j.pmpp.2007.09.003
Kaiser NR, Coombs JJ, Felcher KJ et al (2020) Genome-wide association analysis of common scab resistance and expression profiling of tubers in response to thaxtomin A treatment underscore the complexity of common scab resistance in tetraploid potato. Am J Potato Res 97:513–522. https://doi.org/10.1007/s12230-020-09800-5
Keinath AP, Loria R (1989) Population dynamics of Streptomyces scabies and other actinomycetes as related to common scab of potato. Phytopathology 79:681–687
Klikocka H (2009) Influence of NPK fertilization enriched with S, Mg, and micronutrients contained in liquid fertilizer Insol 7 on potato tubers yield (Solanum tuberosum L.) and infestation of tubers with Streptomyces scabies and Rhizoctonia solani. J Elem 14:271–288
Klikocka H, Haneklaus S, Bloem E, Schnug E (2005) Influence of sulfur fertilization on infection of potato tubers with Rhizoctonia solani and Streptomyces scabies. J Plant Nutr 28:819–833. https://doi.org/10.1081/PLN-200055547
Kobayashi A, Kobayashi YO, Someya N, Ikeda S (2015) Community analysis of root-and tuber-associated bacteria in field-grown potato plants harboring different resistance levels against common scab. Microbes Environ 30:301–309. https://doi.org/10.1264/jsme2.ME15109
Kopecky J, Samkova Z, Sarikhani E et al (2019) Bacterial, archaeal and micro-eukaryotic communities characterize a disease-suppressive or conducive soil and a cultivar resistant or susceptible to common scab. Sci Rep 9:1–14. https://doi.org/10.1038/s41598-019-51570-6
Kopecky J, Rapoport D, Sarikhani E et al (2021) Micronutrients and soil microorganisms in the suppression of potato common scab. Agronomy 11:383. https://doi.org/10.3390/agronomy11020383
Kozich JJ, Westcott SL, Baxter NT et al (2013) Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the MiSeq Illumina sequencing platform. Appl Environ Microbiol 79:5112–5120. https://doi.org/10.1128/AEM.01043-13
Krištůfek V, Diviš J, Omelka M et al (2015) Site, year and cultivar effects on relationships between periderm nutrient contents and common scab severity. Am J Potato Res 92:473–482. https://doi.org/10.1007/s12230-015-9456-6
Lacey MJ, Wilson CR (2001) Relationship of common scab incidence of potatoes grown in Tasmanian ferrosol soils with pH, exchangeable cations and other chemical properties of those soils. J Phytopathol 149:679–683. https://doi.org/10.1046/j.1439-0434.2001.00693.x
Lambert DH, Manzer FE (1991) Relationship of calcium to potato scab. Phytopathology 81:632–636. https://doi.org/10.1094/Phyto-81-632
Lane DJ (1991) 16S/23S rRNA Sequencing. In: Stackebrandt E, Goodfellow M (eds) Nucleic acid techniques in bacterial systematics. Wiley, New York, pp 115–175
Lazarovits G, Hill J, Patterson G et al (2007) Edaphic soil levels of mineral nutrients, pH, organic matter, and cationic exchange capacity in the geocaulosphere associated with potato common scab. Phytopathology 97:1071–1082. https://doi.org/10.1094/PHYTO-97-9-1071
Leiminger J, Frank M, Wenk C et al (2013) Distribution and characterization of Streptomyces species causing potato common scab in Germany. Plant Pathol 62:611–623. https://doi.org/10.1111/j.1365-3059.2012.02659.x
Lerat S, Simao-Beaunoir AM, Beaulieu C (2009) Genetic and physiological determinants of Streptomyces scabies pathogenicity. Mol Plant Pathol 10:579–585. https://doi.org/10.1111/j.1364-3703.2009.00561.x
Lin Y, Ye G, Kuzyakov Y et al (2019) Long-term manure application increases soil organic matter and aggregation, and alters microbial community structure and keystone taxa. Soil Biol Biochem 134:187–196. https://doi.org/10.1016/j.soilbio.2019.03.030
Loria R, Bukhalid RA, Fry BA, King RR (1997) Plant pathogenicity in the genus Streptomyces. Plant Dis 81:836–846. https://doi.org/10.1094/PDIS.1997.81.8.836
Madrova P, Vetrovsky T, Omelka M et al (2018) A short-term response of soil microbial communities to cadmium and organic substrate amendment in long-term contaminated soil by toxic elements. Front Microbiol 9:1–12. https://doi.org/10.3389/fmicb.2018.02807
Manter DK, Delgado JA, Holm DG, Stong RA (2010) Pyrosequencing reveals a highly diverse and cultivar-specific bacterial endophyte community in potato roots. Microb Ecol 60:157–166. https://doi.org/10.1007/s00248-010-9658-x
Martin AP (2002) Phylogenetic approaches for describing and comparing the diversity of microbial communities. Appl Environ Microbiol 68:3673–3682. https://doi.org/10.1128/AEM.68.8.3673
Mendes R, Kruijt M, De Bruijn I et al (2011) Deciphering the rhizosphere microbiome for disease-suppressive bacteria. Science 332:1097–1100. https://doi.org/10.1126/science.1203980
Muyzer G, De Waal EC, Uitterlinden AG (1993) Profiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA. Appl Environ Microbiol 59:695–700
Nahar K, Goyer C, Zebarth BJ et al (2018) Pathogenic Streptomyces spp. abundance affected by potato cultivars. Phytopathology 108:1046–1055. https://doi.org/10.1094/PHYTO-03-18-0075-R
Nakamura K, Hiraishi A, Yoshimi Y et al (1995) Microlunatus phosphovorus gen. nov., sp. nov., a new gram-positive polyphosphate-accumulating bacterium isolated from activated sludge. Int J Syst Bacteriol 45:17–22
Neeno-Eckwall EC, Kinkel LL, Schottel JL (2001) Competition and antibiosis in the biological control of potato scab. Can J Microbiol 47:332–340. https://doi.org/10.1139/w01-010
Oksanen J, Blanchet FG, Friendly M et al (2019) Package “vegan” Title Community Ecology Package Version 2.5-6
Ondov BD, Bergman NH, Phillippy AM (2011) Interactive metagenomic visualization in a Web browser. BMC Bioinformatics 12:385. https://doi.org/10.1186/1471-2105-12-385
Pascale A, Proietti S, Pantelides IS, Stringlis IA (2020) Modulation of the root microbiome by plant molecules: The basis for targeted disease suppression and plant growth promotion. Front Plant Sci 10:1–23. https://doi.org/10.3389/fpls.2019.01741
Qu X, Wanner LA, Christ BJ (2008) Using the txtAB operon to quantify pathogenic Streptomyces in potato tubers and soil. Phytopathology 98:405–412. https://doi.org/10.1094/PHYTO-98-4-0405
R Core Team (2020) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna. https://www.r-project.org. Accessed Oct 2020
Rosenzweig N, Tiedje JM, Quensen JF et al (2012) Microbial communities associated with potato common scab-suppressive soil determined by pyrosequencing analyses. Plant Dis 96:718–725. https://doi.org/10.1094/PDIS-07-11-0571
Rousk J, Bååth E, Brookes PC et al (2010) Soil bacterial and fungal communities across a pH gradient in an arable soil. ISME J 4:1340–1351. https://doi.org/10.1038/ismej.2010.58
Sagova-Mareckova M, Cermak L, Novotna J et al (2008) Innovative methods for soil DNA purification tested in soils with widely differing characteristics. Appl Environ Microbiol 74:2902–2907. https://doi.org/10.1128/AEM.02161-07
Sagova-Mareckova M, Daniel O, Omelka M et al (2015) Determination of factors associated with natural soil suppressivity to potato common scab. PLoS One 10:1–13. https://doi.org/10.1371/journal.pone.0116291
Sagova-Mareckova M, Omelka M, Kopecky J (2017) Sequential analysis of soil factors related to common scab of potatoes. FEMS Microbiol Ecol 93:fiw201. https://doi.org/10.1093/femsec/fiw201
Sarikhani E, Sagova-Mareckova M, Omelka M, Kopecky J (2017) The effect of peat and iron supplements on the severity of potato common scab and bacterial community in tuberosphere soil. FEMS Microbiol Ecol 93:fiw206. https://doi.org/10.1093/femsec/fiw206
Sasse J, Martinoia E, Northen T (2018) Feed your friends: Do plant exudates shape the root microbiome? Trends Plant Sci 23:25–41. https://doi.org/10.1016/j.tplants.2017.09.003
Schlatter D, Kinkel L, Thomashow L et al (2017) Disease suppressive soils: New insights from the soil microbiome. Phytopathology 107:1284–1297. https://doi.org/10.1094/PHYTO-03-17-0111-RVW
Schloss PD, Westcott SL, Ryabin T et al (2009) Introducing mothur: Open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75:7537–7541. https://doi.org/10.1128/AEM.01541-09
Sedláková V, Dejmalová J, Doležal P et al (2013) Characterization of forty-four potato varieties for resistance to common scab, black scurf and silver scurf. Crop Prot 48:82–87. https://doi.org/10.1016/j.cropro.2013.02.014
Shi W, Li M, Wei G et al (2019) The occurrence of potato common scab correlates with the community composition and function of the geocaulosphere soil microbiome. Microbiome 7:1–18. https://doi.org/10.1186/s40168-019-0629-2
Stach JEM, Maldonado LA, Ward AC et al (2003) New primers for the class Actinobacteria: Application to marine and terrestrial environments. Environ Microbiol 5:828–841. https://doi.org/10.1046/j.1462-2920.2003.00483.x
Tokala RK, Strap JL, Jung CM et al (2002) Novel plant-microbe rhizosphere interaction involving Streptomyces lydicus WYEC108 and the pea plant (Pisum sativum). Appl Environ Microbiol 68:2161–2171. https://doi.org/10.1128/AEM.68.5.2161-2171.2002
Trujillo ME, Alonso-Vega P, Rodríguez R et al (2010) The genus Micromonospora is widespread in legume root nodules: The example of Lupinus angustifolius. ISME J 4:1265–1281. https://doi.org/10.1038/ismej.2010.55
Wang A, Lazarovits G (2005) Role of seed tubers in the spread of plant pathogenic Streptomyces and initiating potato common scab disease. Am J Potato Res 82:221–230. https://doi.org/10.1007/BF02853588
Wanner LA (2009) A patchwork of Streptomyces species isolated from potato common scab lesions in North America. Am J Potato Res 86:247–264. https://doi.org/10.1007/s12230-009-9078-y
Wenzl H, Demel J (1967) Bildskalen für die Beurteilung von Kartoffelschorf und Rhizoctonia-Pocken. Der Pflanzenarzt 20:77–78
White JR, Nagarajan N, Pop M (2009) Statistical methods for detecting differentially abundant features in clinical metagenomic samples. PLoS Comput Biol 5:e1000352. https://doi.org/10.1371/journal.pcbi.1000352
Xue D, Christenson R, Genger R et al (2018) Soil microbial communities reflect both inherent soil properties and management practices in Wisconsin potato fields. Am J Potato Res 95:696–708. https://doi.org/10.1007/s12230-018-9677-6
Yamada T, Hamada M, Nakagawa M et al (2019) 16S rRNA gene amplicon sequencing of microbiota in polybutylene succinate adipate-packed denitrification reactors used for water treatment of land-based recirculating aquaculture systems. Microbiol Resour Announc 8:14–16. https://doi.org/10.1128/mra.01295-19
Yan Y, Kuramae EE, De Hollander M et al (2017) Functional traits dominate the diversity-related selection of bacterial communities in the rhizosphere. ISME J 11:56–66. https://doi.org/10.1038/ismej.2016.108
Yilmaz P, Parfrey LW, Yarza P et al (2014) The SILVA and “All-species Living Tree Project (LTP)” taxonomic frameworks. Nucleic Acids Res 42:D643–D648. https://doi.org/10.1093/nar/gkt1209
Youssef NH, Elshahed MS (2009) Diversity rankings among bacterial lineages in soil. ISME J 3:305–313. https://doi.org/10.1038/ismej.2008.106
Funding
This work was supported by the Ministry of Agriculture of the Czech Republic, project QK1810370 and Institutional Project RO4018.
Author information
Authors and Affiliations
Contributions
M.S-M. organized the results and wrote the MS. E.S. and O.D. extracted DNAs, prepared PCRs. M.O. did statistical analyses. V.K., J.D., M.S-M. and J.K. collected samples at the sites. J.K. analyzed bacterial communities, and created figures and tables.
Corresponding author
Ethics declarations
Conflicts of interest/Competing interests
All authors declare no conflict of interest.
Additional information
Responsible Editor: Hans Lambers.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
(PDF 7.55 MB)
Rights and permissions
About this article
Cite this article
Marketa, SM., Sarikhani, E., Daniel, O. et al. Tuberosphere and bulk soil microbial communities in fields differing in common scab severity are distinguished by soil chemistry and interactions with pathogens. Plant Soil 468, 259–275 (2021). https://doi.org/10.1007/s11104-021-05128-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-021-05128-z