Abstract
Background and aims
The rhizosphere microbiome has been shown to contribute to nutrient acquisition, protection against biotic and abiotic stresses and, ultimately, to changes in the development and physiology of plants. Here, using a controlled natural selection approach, we followed the microbial dynamics in the soil of Arabidopsis thaliana plants infected with the foliar pathogen Pseudomonas syringae DC3000 (Pst).
Methods
Plants were iteratively cultivated on a pasteurised soil inoculated with the soil microbial community of the previous iteration isolated from the rhizosphere of plants infected with Pst (pst-line) or not (mock-line). Modification of soil microbial communities was assessed through an amplicon-based metagenomic analysis targeting bacterial and fungal diversity. Plant fitness and transcript abundance of stress hormone related genes were also analysed.
Results
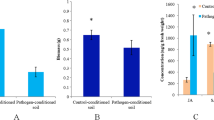
At the tenth and eleventh iterations respectively, we observed a reduction in disease severity of 81% and 85% in pst-lines as compared to mock-lines. These changes were associated with (i) an early induction of defence mechanisms mediated by salicylic acid, in pst-line as compared to mock-line, shown by the decrease in transcript abundance of salicylic acid related genes, whereas jasmonic acid, ethylene or abscisic acid related genes remained unchanged and (ii) a shift in soil bacterial, and not in fungal, composition.
Conclusions
Our study suggests that these changes in soil bacterial composition are mediated by plant-soil feedback in response to Pst and resulted in an activation of SA-related immune response in the plant. This supports the concept of applying plant-soil feedbacks to enhance soil suppressiveness against foliar pathogens.





Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Allen WJ, Sapsford SJ, Dickie IA (2021) Soil sample pooling generates no consistent inference bias: a meta-analysis of 71 plant–soil feedback experiments. New Phytol 231:1308–1315. https://doi.org/10.1111/nph.17455
Bais HP, Weir TL, Perry LG et al (2006) The role of root exudates in rhizosphere interactions with plants and other organisms. Annu Rev Plant Biol 57:233–266. https://doi.org/10.1146/annurev.arplant.57.032905.105159
Berendsen RL, Pieterse CMJ, Bakker PAHM (2012) The rhizosphere microbiome and plant health. Trends Plant Sci 17:478–486. https://doi.org/10.1016/j.tplants.2012.04.001
Berens ML, Wolinska KW, Spaepen S et al (2019) Balancing trade-offs between biotic and abiotic stress responses through leaf age-dependent variation in stress hormone cross-talk. Proc Natl Acad Sci U S A 116:2364–2373. https://doi.org/10.1073/pnas.1817233116
Cao M-J, Zhang Y-L, Liu X et al (2017) Combining chemical and genetic approaches to increase drought resistance in plants. Nat Commun 8:1183. https://doi.org/10.1038/s41467-017-01239-3
Chapelle E, Mendes R, Bakker PAHM, Raaijmakers JM (2016) Fungal invasion of the rhizosphere microbiome. The ISME Journal 10:265–268. https://doi.org/10.1038/ismej.2015.82
Conner JK (2003) Artificial selection: a powerful tool for ecologists. Ecology 84:1650–1660
Cordovez V, Dini-Andreote F, Carrión VJ, Raaijmakers JM (2019) Ecology and evolution of plant microbiomes. Annu Rev Microbiol 73:69–88. https://doi.org/10.1146/annurev-micro-090817-062524
Dalal M, Tayal D, Chinnusamy V, Bansal KC (2009) Abiotic stress and ABA-inducible Group 4 LEA from Brassica napus plays a key role in salt and drought tolerance. J Biotechnol 139:137–145. https://doi.org/10.1016/j.jbiotec.2008.09.014
De Vos M, Van Oosten VR, Van Poecke RMP et al (2005) Signal signature and transcriptome changes of Arabidopsis during pathogen and insect attack. Mol Plant-Microbe Interact 18:923–937. https://doi.org/10.1094/MPMI-18-0923
Dodd ICC, Zinovkina NYY, Safronova VII, Belimov AAA (2010) Rhizobacterial mediation of plant hormone status. Ann Appl Biol 157:361–379. https://doi.org/10.1111/j.1744-7348.2010.00439.x
Doornbos R, van Loon L, Bakker P (2012) Impact of root exudates and plant defense signaling on bacterial communities in the rhizosphere. A review Agronomy for Sustainable Development 32:227–243. https://doi.org/10.1007/s13593-011-0028-y
Gardes M, Bruns TD (1993) ITS primers with enhanced specificity for basidiomycetes - application to the identification of mycorrhizae and rusts. Mol Ecol 2:113–118. https://doi.org/10.1111/j.1365-294X.1993.tb00005.x
Griffiths RI, Whiteley AS, O’Donnell AG, Bailey MJ (2000) Rapid method for coextraction of DNA and RNA from natural environments for analysis of ribosomal DNA- and rRNA-based microbial community composition. Appl Environ Microbiol 66:5488–5491. https://doi.org/10.1128/AEM.66.12.5488-5491.2000
Hase S, Van Pelt JA, Van Loon LC, Pieterse CMJ (2003) Colonization of Arabidopsis roots by Pseudomonas fluorescens primes the plant to produce higher levels of ethylene upon pathogen infection. Physiol Mol Plant Pathol 62:219–226. https://doi.org/10.1016/S0885-5765(03)00059-6
Jacquiod S, Spor A, Wei S et al (2022) Artificial selection of stable rhizosphere microbiota leads to heritable plant phenotype changes. Ecol Lett 25:189–201. https://doi.org/10.1111/ele.13916
Janda M, Ruelland E (2015) Magical mystery tour: salicylic acid signalling. Environ Exp Bot 114:117–128. https://doi.org/10.1016/j.envexpbot.2014.07.003
Katagiri F, Thilmony R, He SY (2002) The Arabidopsis Thaliana-pseudomonas Syringae interaction. The Arabidopsis Book 1:e0039. https://doi.org/10.1199/tab.0039
Klessig DF, Choi HW, Dempsey DA (2018) Systemic acquired resistance and salicylic acid: past, present, and future. Mol Plant-Microbe Interact 31:871–888. https://doi.org/10.1094/MPMI-03-18-0067-CR
Klindworth A, Pruesse E, Schweer T et al (2013) Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res 41:1–11. https://doi.org/10.1093/nar/gks808
Kulmatiski A, Beard KH, Stevens JR, Cobbold SM (2008) Plant–soil feedbacks: a meta-analytical review. Ecol Lett 11:980–992. https://doi.org/10.1111/j.1461-0248.2008.01209.x
Lemanceau P, Blouin M, Muller D, Moënne-Loccoz Y (2017) Let the Core microbiota be functional. Trends Plant Sci. https://doi.org/10.1016/j.tplants.2017.04.008
Leon-Reyes A, Van der Does D, De Lange ES et al (2010) Salicylate-mediated suppression of jasmonate-responsive gene expression in Arabidopsis is targeted downstream of the jasmonate biosynthesis pathway. Planta 232:1423–1432. https://doi.org/10.1007/s00425-010-1265-z
Leontovyčová H, Kalachova T, Trdá L et al (2019) Actin depolymerization is able to increase plant resistance against pathogens via activation of salicylic acid signalling pathway. Sci Rep 9:1–10. https://doi.org/10.1038/s41598-019-46465-5
Liu L, Sonbol F-M, Huot B et al (2016) Salicylic acid receptors activate jasmonic acid signalling through a non-canonical pathway to promote effector-triggered immunity. Nat Commun 7:13099. https://doi.org/10.1038/ncomms13099
Luna E, Bruce TJA, Roberts MR et al (2012) Next-generation systemic acquired resistance. Plant Physiol 158:844–853. https://doi.org/10.1104/pp.111.187468
Mahé F, Rognes T, Quince C et al (2014) Swarm: robust and fast clustering method for amplicon-based studies. PeerJ 2014:1–13. https://doi.org/10.7717/peerj.593
McMurdie PJ, Holmes S (2013) Phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS One 8. https://doi.org/10.1371/journal.pone.0061217
Meaden S, Koskella B (2017) Adaptation of the pathogen, pseudomonas syringae, during experimental evolution on a native vs. alternative host plant. Mol Ecol 26:1790–1801. https://doi.org/10.1111/mec.14060
Mendes R, Kruijt M, de Bruijn I et al (2011) Deciphering the rhizosphere microbiome for disease-suppressive bacteria. Science (New York, NY) 332:1097–1100. https://doi.org/10.1126/science.1203980
Mitter B, Brader G, Afzal M, et al (2013) Advances in Elucidating Beneficial Interactions Between Plants, Soil, and Bacteria. In: Sparks DLBT-A in A (ed). Academic Press, pp. 381–445
Monard C, Gantner S, Stenlid J (2013) Utilizing ITS1 and ITS2 to study environmental fungal diversity using pyrosequencing. FEMS Microbiol Ecol 84:165–175. https://doi.org/10.1111/1574-6941.12046
Mueller UG, Juenger TE, Kardish MR et al (2021) Artificial selection on microbiomes to breed microbiomes that confer salt tolerance to plants. mSystems 6:e01125–e01121. https://doi.org/10.1128/mSystems.01125-21
Nicolaisen MH, Bælum J, Jacobsen CS, Sørensen J (2008) Transcription dynamics of the functional tfdA gene during MCPA herbicide degradation by Cupriavidus necator AEO106 (pRO101) in agricultural soil. Environ Microbiol 10:571–579. https://doi.org/10.1111/j.1462-2920.2007.01476.x
Niu D-D, Liu H-X, Jiang C-H et al (2011) The plant growth–promoting Rhizobacterium Bacillus cereus AR156 induces systemic resistance in Arabidopsis thaliana by simultaneously activating Salicylate- and Jasmonate/ethylene-dependent signaling pathways. Molecular plant-microbe interactions® 24:533–542. https://doi.org/10.1094/MPMI-09-10-0213
Panke-Buisse K, Poole AC, Goodrich JK et al (2015) Selection on soil microbiomes reveals reproducible impacts on plant function. The ISME journal 9:980–989. https://doi.org/10.1038/ismej.2014.196
Pathma J, Sakthivel N (2013) Molecular and functional characterization of bacteria isolated from straw and goat manure based vermicompost. Appl Soil Ecol 70:33–47. https://doi.org/10.1016/j.apsoil.2013.03.011
Pieterse CMJ, Leon-Reyes A, Van der Ent S, Van Wees SCM (2009) Networking by small-molecule hormones in plant immunity. Nat Chem Biol 5:308–316. https://doi.org/10.1038/nchembio.164
Pieterse CMJ, de Jonge R, Berendsen RL (2016) The soil-borne supremacy. Trends Plant Sci 21:171–173. https://doi.org/10.1016/j.tplants.2016.01.018
Pineda A, Kaplan I, Bezemer TM (2017) Steering soil microbiomes to suppress aboveground insect pests. Trends Plant Sci 22:770–778. https://doi.org/10.1016/j.tplants.2017.07.002
Pineda A, Kaplan I, Hannula SE et al (2020) Conditioning the soil microbiome through plant–soil feedbacks suppresses an aboveground insect pest. New Phytol 226:595–608. https://doi.org/10.1111/nph.16385
Puga-Freitas R, Blouin M (2015) A review of the effects of soil organisms on plant hormone signalling pathways. Environ Exp Bot 114. https://doi.org/10.1016/j.envexpbot.2014.07.006
Qi G, Chen J, Chang M et al (2018) Pandemonium breaks out: disruption of salicylic acid-mediated defense by plant pathogens. Mol Plant 11:1427–1439. https://doi.org/10.1016/j.molp.2018.10.002
R Core Team (2020) R: A Language and Environment for Statistical Computing. https://www.r-project.org/. Vienna, Austria
Raaijmakers JM, Mazzola M (2016) Soil immune responses. Science 352:1392–1393. https://doi.org/10.1126/science.aaf3252
Raaijmakers J, Paulitz T, Steinberg C et al (2009) The rhizosphere: a playground and battlefield for soilborne pathogens and beneficial microorganisms. Plant Soil 321:341–361. https://doi.org/10.1007/s11104-008-9568-6
Robert-Seilaniantz A, Grant M, Jones JDG (2011) Hormone crosstalk in plant disease and defense: more than just JASMONATE-SALICYLATE antagonism. Annu Rev Phytopathol 49:317–343. https://doi.org/10.1146/annurev-phyto-073009-114447
Rolfe SA, Griffiths J, Ton J (2019) Crying out for help with root exudates: adaptive mechanisms by which stressed plants assemble health-promoting soil microbiomes. Curr Opin Microbiol 49:73–82. https://doi.org/10.1016/j.mib.2019.10.003
Rudrappa T, Czymmek KJ, Paré PW, Bais HP (2008) Root-secreted malic acid recruits beneficial soil Bacteria. Plant Physiol 148:1547–1556. https://doi.org/10.1104/pp.108.127613
Rymaszewski W, Dauzat M, Bédiée A et al (2018) Measurement of Arabidopsis thaliana plant traits using the PHENOPSIS phenotyping platform. Bio-protocol 8:e2739. https://doi.org/10.21769/BioProtoc.2739
Sanguin H, Sarniguet A, Gazengel K et al (2009) Rhizosphere bacterial communities associated with disease suppressiveness stages of take-all decline in wheat monoculture. New Phytol 184:694–707. https://doi.org/10.1111/j.1469-8137.2009.03010.x
Schlatter D, Kinkel L, Thomashow L et al (2017) Disease suppressive soils: new insights from the soil microbiome. Phytopathology® 107:1284–1297. https://doi.org/10.1094/PHYTO-03-17-0111-RVW
Semchenko M, Barry KE, de Vries FT et al (2022) Deciphering the role of specialist and generalist plant–microbial interactions as drivers of plant–soil feedback. New Phytol 234:1929–1944. https://doi.org/10.1111/nph.18118
Steiner AA (1961) A universal method for preparing nutrient solutions of a certain desired composition. Plant Soil 15:134–154. https://doi.org/10.1007/BF01347224
Su F, Villaume S, Rabenoelina F et al (2017) Different Arabidopsis thaliana photosynthetic and defense responses to hemibiotrophic pathogen induced by local or distal inoculation of Burkholderia phytofirmans. Photosynth Res 134:201–214. https://doi.org/10.1007/s11120-017-0435-2
Swenson W, Wilson DS, Elias R (2000) Artificial ecosystem selection. Proc Natl Acad Sci U S A 97:9110–9114. https://doi.org/10.1073/pnas.150237597
Teste FP, Paul K, Turner BL et al (2017) Plant-soil feedback and the maintenance of diversity in Mediterranean-climate shrublands. Science 355:173–176. https://doi.org/10.1126/science.aai8291
Thioulouse J, Dray S, Dufour A et al (2018) Multivariate analysis of ecological data with ade4. Springer. https://doi.org/10.1007/978-1-4939-8850-1
Trivedi P, Mattupalli C, Eversole K, Leach JE (2021) Enabling sustainable agriculture through understanding and enhancement of microbiomes. New Phytol. https://doi.org/10.1111/nph.17319
van der Putten WH, Bardgett RD, Bever JD et al (2013) Plant–soil feedbacks: the past, the present and future challenges. J Ecol 101:265–276. https://doi.org/10.1111/1365-2745.12054
Wang KL-C, Li H, Ecker JR (2002) Ethylene biosynthesis and signaling networks. Plant Cell 14:S131–S151. https://doi.org/10.1105/tpc.001768
Weller DM, Raaijmakers JM, Gardener BBM, Thomashow LS (2002) Microbial populations responsible for specific soil suppressiveness to plant pathogens. Annu Rev Phytopathol 40:309–348. https://doi.org/10.1146/annurev.phyto.40.030402.110010
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJBT-PCRP (eds) PCR protocols: a guide to methods and applications. Academic Press, San Diego, pp 315–322
Yang L, Li B, Zheng X et al (2015) Salicylic acid biosynthesis is enhanced and contributes to increased biotrophic pathogen resistance in Arabidopsis hybrids. Nat Commun 6:7309. https://doi.org/10.1038/ncomms8309
Yuan J, Zhao J, Wen T et al (2018) Root exudates drive the soil-borne legacy of aboveground pathogen infection. Microbiome 6:156. https://doi.org/10.1186/s40168-018-0537-x
Acknowledgements
We thank Sophie Michon-Coudouel and Romain Causse-Védrine from the EcogenO Platform (OSUR, France) for the preparation of the sequençing libraries and the sequencing. We are also grateful to the genotoul bioinformatics platform Toulouse Occitanie (Bioinfo Genotoul, https://doi.org/10.15454/1.5572369328961167E12) for providing computing resources, to Dr. Aleš Hanč from the Czech University of Life Sciences (Prague), for providing the vermicompost and to Pr. Anne Repellin from iEES-Paris, for English language editing and comments on the manuscript.
Funding
ER and RPF benefitted from an EC2CO initiative funding from CNRS. This work was also supported by the European Regional Development Fund, Project “Centre for Experimental Plant Biology” [grant no. CZ.02.1.01/0.0/0.0/16_019/0000738] and grant from the Contact Mobility - Barrande Program (PHC action) awarded to LB and RPF.
Author information
Authors and Affiliations
Contributions
TK, LB, ER and RPF conceived and designed the study. TK and BJ performed the experiment. CM performed the soil DNA extraction and sequencing. RPF analysed the data. TK, MB, SJ, ER and RPF interpreted the data. TK, ER and RPF wrote the first manuscript and all authors contributed to the revision of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Responsible Editor: Jonathan Richard De Long.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
(DOCX 822 kb)
Rights and permissions
About this article
Cite this article
Kalachova, T., Jindřichová, B., Burketová, L. et al. Controlled natural selection of soil microbiome through plant-soil feedback confers resistance to a foliar pathogen. Plant Soil 485, 181–195 (2023). https://doi.org/10.1007/s11104-022-05597-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-022-05597-w