-
PDF
- Split View
-
Views
-
Cite
Cite
Lucia Tomečková, Aleš Tomčala, Miroslav Oborník, Vladimír Hampl, The Lipid Composition of Euglena gracilis Middle Plastid Membrane Resembles That of Primary Plastid Envelopes, Plant Physiology, Volume 184, Issue 4, December 2020, Pages 2052–2063, https://doi.org/10.1104/pp.20.00505
Close - Share Icon Share
Abstract
Euglena gracilis is a photosynthetic flagellate possessing chlorophyte-derived secondary plastids that are enclosed by only three enveloping membranes, unlike most secondary plastids, which are surrounded by four membranes. It has generally been assumed that the two innermost E. gracilis plastid envelopes originated from the primary plastid, while the outermost is of eukaryotic origin. It was suggested that nucleus-encoded plastid proteins pass through the middle and innermost plastid envelopes of E. gracilis by machinery homologous to the translocons of outer and inner chloroplast membranes, respectively. Although recent genomic, transcriptomic, and proteomic data proved the presence of a reduced form of the translocon of inner membrane, they failed to identify any outer-membrane translocon homologs, which raised the question of the origin of E. gracilis’s middle plastid envelope. Here, we compared the lipid composition of whole cells of the pigmented E. gracilis strain Z and two bleached mutants that lack detectable plastid structures, W10BSmL and WgmZOflL. We determined the lipid composition of E. gracilis strain Z mitochondria and plastids, and of plastid subfractions (thylakoids and envelopes), using HPLC high-resolution tandem mass spectrometry, thin-layer chromatography, and gas chromatography-flame ionization detection analytical techniques. Phosphoglycerolipids are the main structural lipids in mitochondria, while glycosyldiacylglycerols are the major structural lipids of plastids and also predominate in extracts of whole mixotrophic cells. Glycosyldiacylglycerols were detected in both bleached mutants, indicating that mutant cells retain some plastid remnants. Additionally, we discuss the origin of the E. gracilis middle plastid envelope based on the lipid composition of envelope fraction.
Euglena gracilis is a freshwater photosynthetic flagellate belonging to the phylum Euglenozoa (Excavata) together with other free-living euglenids, free-living and parasitic kinetoplastids, planktonic diplonemids, and epibiont-carrying symbiontids (Adl et al., 2012, 2019). E. gracilis, like other euglenophytes, possesses secondary plastids that are descendants of a green alga, of which the closest extant relative is the marine prasinophyte Pyramimonas parkeae (Turmel et al., 2009; Hrdá et al., 2012; Pombert et al., 2012; Wiegert et al., 2012, 2013; Bennett et al., 2014; Bennett and Triemer, 2015; Dabbagh and Preisfeld, 2017).
Most secondary plastids are surrounded by four envelopes: the two innermost are derived from the primary plastid and originally from a cyanobacterium, while the two outermost are of eukaryotic origin, derived from the plasma membrane of the primary alga (the third) and phagosome of the host (the fourth). Plastids of euglenophytes and peridinin-containing dinoflagellates are, however, enclosed by only three envelopes, suggesting that either one of the envelopes has been lost or the mechanism of acquisition of these plastids led to only three membranes (Gibbs, 1978; Lefort‐Tran, 1981). The latter can be achieved by myzocytosis (i.e. by perforation of the plasma membrane and sucking the organelles from the cytoplasm; Schnepf and Deichgräber, 1984).
The presence of additional envelopes necessitates a more complex protein import system for secondary plastids. In E. gracilis, plastid-targeted nucleus-encoded precursor proteins contain N-terminal signal peptides, which are recognized by the signal recognition particle. Precursor proteins either are cotranslationally transported into the lumen of the endoplasmic reticulum (class II) or stay anchored in the endoplasmic reticulum membrane (class I; Durnford and Gray, 2006). Following signal peptide cleavage, a transit peptide is revealed, guiding the protein’s transport through the Golgi apparatus (Sulli and Schwartzbach, 1995, 1996; Sulli et al., 1999; van Dooren et al., 2001). Transport vesicles carrying plastid precursor proteins then fuse with the outermost plastid envelope by an unknown mechanism, and the proteins are released to the space beyond this membrane. The fact that the outermost euglenophyte plastid membrane can fuse with Golgi-derived vesicles carrying plastid precursor proteins (Sulli et al., 1999) suggests that it is derived from the phagosomal membrane of euglenids (Cavalier-Smith, 1999, 2003).
Based on the presence of the transit peptide, it was suggested that proteins are further transported by machinery homologous to the primary plastid translocons of the outer (TOC) and inner (TIC) chloroplast membranes, presumably residing in the middle and the innermost plastid envelopes of euglenophytes, respectively (Inagaki et al., 2000; Sláviková et al., 2005). Recent transcriptomic and proteomic studies indeed identified a reduced form of the TIC complex but failed to identify either TOC homologs (Záhonová et al., 2018; Ebenezer et al., 2019; Novák Vanclová et al., 2020) or any representative of porin family proteins specific to the outer membrane of primary plastids. Instead, the proteomic study (Novák Vanclová et al., 2020) reported the presence of putative derlin-like pseudoproteases very distantly related to those involved in SELMA (symbiont-derived ERAD-like machinery), which facilitate protein transport across the third membrane of several secondary plastids. This complex is derived from the endoplasmic reticulum-associated degradation (ERAD) pathway and is of eukaryotic origin. Although localization of these proteins is not known, their presence casts doubt on the assumption that the middle envelope of E. gracilis plastids is derived from the primary plastid and raises the possibility that the primary plastid outer membrane could be absent.
Glycosyldiacylglycerols (GDAGs) are signature lipids of all photosynthetic eukaryotes. They are considered to be the most abundant lipid class on Earth, constituting more than 70% of all structural lipids in the cells of primary algae, plants, and cyanobacteria (Gounaris and Barber, 1983). They contain one (monogalactosyldiacylglycerol [MGDG]), two (digalactosyldiacylglycerol [DGDG]), or more molecules of Gal or sulfoquinovose (sulfoquinovosyldiacylglycerol [SQDG]) bound to a glycerol backbone carrying two fatty acid (FA) chains. Neutrally charged MGDGs and DGDGs, together with negatively charged SQDGs and phosphatidylglycerols (PGs), form the thylakoids and plastid envelopes of primary plastids and harbor photosystems. On the other hand, phosphatidylcholines (PCs) and phosphatidylethanolamines (PEs) are considered to be typical extrathylakoid lipids and major constituents of the other membranes in the cells of phototrophs (Harwood, 1998).
Most of the studies concerning lipid and FA composition and their synthesis in plastids were first performed on the primary plastids of Arabidopsis (Arabidopsis thaliana) or spinach (Spinacia oleracea). Under standard conditions, the synthesis of MGDGs occurs at the inner plastid envelope membrane (Miège et al., 1999; Awai et al., 2001; for review, see Petroutsos et al., 2014; LaBrant et al., 2018), where MGDG synthase (MGD1) transfers Gal from UDP-Gal to diacylglycerol and forms MGDGs. DGDG synthase (DGD1), operating at the outer plastid envelope, adds a second Gal to MGDGs and produces DGDGs (Froehlich et al., 2001; for review, see Petroutsos et al., 2014; LaBrant et al., 2018). The synthesis of SQDGs starts with the synthesis of sulfoquinovose, which serves as the sulfolipid head group donor. The reaction is typically catalyzed by the UDP-sulfoquinovose synthase (SQD1; Sanda et al., 2001; for review, see Petroutsos et al., 2014; LaBrant et al., 2018), which is absent in E. gracilis, and so an alternative mechanism is assumed there (Novák Vanclová et al., 2020). The second step of SQDG synthesis is performed by sulfoquinovosyltransferase (SQD2) and is localized at the inner plastid envelope (Yu et al., 2002; Frentzen, 2004; Simm et al., 2013).
While the outer primary plastid envelope is enriched with DGDGs and PCs (Dorne et al., 1985), the inner envelope and thylakoids are enriched in MGDGs and PGs. SQDGs are equally distributed in all primary plastid fractions and, along with PGs, provide for the correct function of photosynthetic membranes, especially in phosphate-limited environments (Yu et al., 2002; Frentzen, 2004). The membranes of other organelles, such as mitochondria and peroxisomes, are composed strictly of phospholipids (PLs), but during phosphorus deprivation, DGDGs synthesized by DGD2 are transported through membrane contacts to the mitochondria to compensate for the lack of PLs in the mitochondrial membranes (Jouhet et al., 2004; for review, see Block et al., 2007).
Lipid and FA composition has been studied in E. gracilis grown under different environmental parameters, including varying temperature (Craig et al., 2015) and heterotrophic, autotrophic, mixotrophic (Hulanicka et al., 1964; Rosenberg et al., 1966; Matson et al., 1970; Pohl and Wagner, 1972; Regnault et al., 1995; Shibata et al., 2018; Wang et al., 2018) and anaerobic (Ogawa et al., 2014) growth, with all studies confirming the presence of GDAGs in E. gracilis cells. A relationship between the lipid content of developing plastids and mitochondria of E. gracilis was recently described (Shibata et al., 2018). The identification of genes encoding GDAG synthesis enzymes in E. gracilis transcriptomic and proteomic data (Yoshida et al., 2016; Shibata et al., 2018; Novák Vanclová et al., 2020) corroborates the biochemical data.
Here, we analyze the lipid composition of a mixotrophic strain, E. gracilis Z, and two bleached heterotrophic mutants, W10BSmL (W10) and WgmZOflL (OFL), obtained by treatment of the E. gracilis bacillaris strain with the antibiotic streptomycin and by an ofloxacin treatment of E. gracilis strain Z, respectively, resulting in both mutant strains lacking detectable plastid structures (Osafune and Schiff, 1983; Polónyi et al., 1998). We further identify the structural lipids in plastid and mitochondrial fractions as well as in the subfractions of plastids: thylakoids and envelopes.
RESULTS
Lipid Composition of Whole Cells of E. gracilis Strains Indicates That Bleached Mutants Contain Plastid Remnants
We analyzed the lipid composition of the pigmented E. gracilis strain Z and the bleached strains OFL and W10 with several analytical techniques: thin-layer chromatography (TLC), gas chromatography-flame ionization detection (GC-FID), and high-performance LC (HPLC) high-resolution tandem mass spectrometry (HR-MS/MS). Four biological replicates were measured for each E. gracilis strain. The data set consisting of peak areas of particular intact lipid species obtained by HPLC HR-MS/MS was processed by correspondence analysis (CA; Supplemental Fig. S1). The subsequent constrained correspondence analysis (CCA) and Monte-Carlo permutation test revealed significant differences among E. gracilis strains (P = 0.001). Since the x axis explains 84.7% of the variability, the 2D spatial distribution of samples directly shows that lipidomic profiles from both bleached strains are more similar to each other than to the mixotrophic E. gracilis. This result was expected because the mutants lost most of their plastid membranes during the bleaching. The differences between W10 and OFL mutants can be attributed to the fact that they were established from different E. gracilis strains and by different bleaching agents. To demonstrate these differences in detail, we compared the abundance of 11 representative lipid species (Fig. 1A), coupled (Fig. 1B) and individual (Fig. 1C) lipid classes, and FAs (Fig. 1D). All these analyses focused only on structural lipids; storage lipids, such as triacylglycerols and waxes, were excluded for clarity. Twenty-two FAs were detected in total, forming 78 lipid species in eight lipid classes (Supplemental Tables S1 and S2).
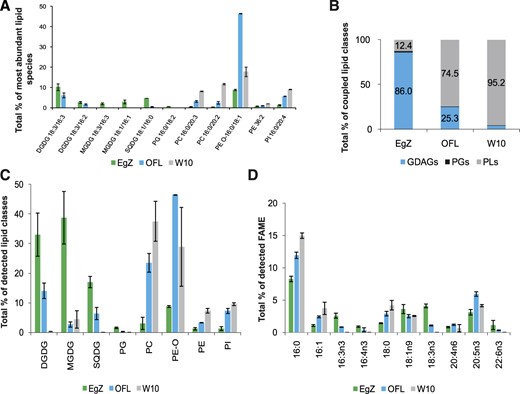
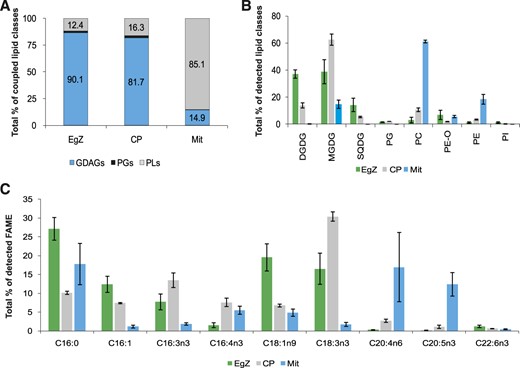
Lipid and FA composition of E. gracilis strain Z (EgZ) and two bleached mutants, OFL and W10. The data were obtained by HPLC HR-MS/MS, and four biological replicates were measured for each E. gracilis strain. sd values are expressed as error bars. A, Graph of chosen intact lipid species in E. gracilis strains. Depicted lipid species represent approximately 35%, 67%, and 49% of all detected lipids in E. gracilis strains Z, OFL, and W10, respectively. B, Bar graph showing percentages of the coupled lipid classes. GDAGs include DGDGs, MGDGs, and SQDGs; PLs include PCs, PE-Os, PEs, and PIs. C, Bar graph showing percentages of detected lipid classes. Data were obtained by coupling peak areas of intact lipid species. D, Bar graph showing percentages of selected fatty acid methyl esters (FAMEs) detected by GC-FID analyses (n = 4). Depicted FAs represent approximately 30% of all detected FAMEs in all EgZ, OFL, and W10 strains.
Figure 1C represents the percentage of all structural lipid classes detected by means of HPLC HR-MS/MS. All membranes of E. gracilis strain Z are composed of eight classes of glycerolipids, mainly represented by GDAGs (MGDGs, DGDGs, and SQDGs), constituting more than 85% of all structural lipids in the cells (Fig. 1B). Interestingly, the most abundant glycerolipid classes in E. gracilis bleached mutants OFL and W10 are plasmanyl phosphatidylethanolamines (PE-Os), in which one FA is linked by an ether bond and the second FA is linked by an ester bond to the glycerol backbone (Hsu and Turk, 2007). PE-Os are represented only by two compounds, PE O-16:0/18:1 and PE O-16:0/18:2, where the former constitutes more than 90% of the total PE-O content. The second most abundant class in these strains are PCs (Fig. 1C). PE-Os and PCs constitute more than 65% of the detected lipids in both bleached strains. Furthermore, the level of another PL class, phosphatidylinositols (PIs), is substantially higher in the bleached strains than in mixotrophic E. gracilis. The lipid species composition of the W10 mutant is different from that of the OFL mutant, which possesses more GDAGs and higher levels of PE-Os (Fig. 1, B and C). GDAGs constitute more than 25% of all structural lipids in OFL mutant cells, while in W10 mutants it is less than 5% (Fig. 1B). Moreover, SQDGs are completely missing in W10 mutants (Fig. 1C), and PGs are present only in trace amounts. GDAGs in bleached OFL and W10 E. gracilis mutants suggest that at least some remnants of plastids persist.
The most abundant lipid species in E. gracilis strain Z are DGDGs and MGDGs, with an α-linolenic FA (C18:3n3) in the sn-2 position and a cis-7,10,13-hexadecatrienoic acid (C16:3n3) in the sn-1 position, followed by a PE-O with palmitic acid (C16:0) as the alkyl in the sn-1 position and oleic acid (C18:1n9) in the sn-2 position as the acyl, PE O-16:0/18:1 (Fig. 1A). It was not possible to detect the positions of double bonds in FAs by using the HPLC HR-MS/MS methodology; therefore, the GC-FID technique was employed. The most abundant lipid species in both bleached mutants (OFL and W10) was again PE O-16:0/18:1, followed by PC 16:0/20:2 and PI 16:0/20:4 (Fig. 1A). The graph in Figure 1A also demonstrates the richness of lipid species in strain Z in comparison with OFL and W10 mutants. The five most abundant GDAGs from strain Z combined did not represent more than 36% of all detected lipids, while the five most abundant PL species constitute 67% and 49% of all OFL and W10 lipids, respectively.
The composition of FAs was also analyzed by GC-FID methodology, and in total, 22 FAs were detected. For clarity, the FAs are shown according to their importance (Fig. 1D). The shorter polyunsaturated fatty acids (PUFAs; C16:3n3, C16:4n3, and C18:3n3) constitute GDAGs and PGs (Supplemental Table S1) exclusively. The quantities of GDAGs and PGs in both mutants were substantially lower than those in the mixotrophic strain, compared to which the amounts of these FAs were less than half in the OFL strain and almost zero in W10 (Fig. 1D). Contrarily, saturated and monounsaturated FAs occur mostly in PLs; note, for instance, the huge amount of PE O-16:0/18:1 (Supplemental Table S2). Interestingly, the value recorded for eicosapentanoic acid (20:5n3) is comparable across all three E. gracilis strains (Supplemental Table S2). However, no docosahexaenoic acid (22:6n3) is present in the W10 mutant (Fig. 1D). Since GDAGs with PUFAs are the main compounds in plastids, all three results perfectly reflect the physiology of these E. gracilis strains.
Preparation of Cell Fractions and Purity Assessment
The lipid composition of the fractions of mitochondria and plastids, and that of the two plastid subfractions, thylakoid and envelope membranes, were analyzed for E. gracilis strain Z. Three biological replicates of each fraction were prepared. Initially, the purity of these fractions was assessed by transmission electron microscopy (TEM) and by a chemical analytical approach.
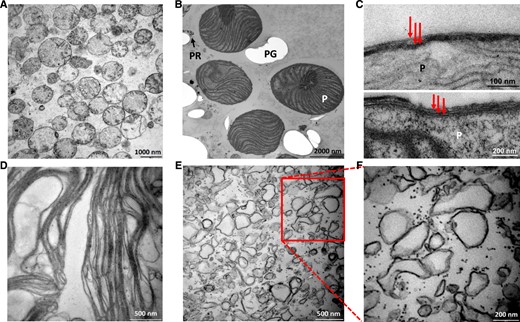
Electron micrographs of these fractions are presented in Figure 2. Mitochondrial fractions (Fig. 2A) consist of mitochondria with visible cristae. No obvious contamination with other cellular compartments was observed. Plastid fractions (Fig. 2B) contained intact plastids, in some of which were visible three envelope membranes (Fig. 2C), holes in the resin after paramylon grains, and a small amount of pellicle residue. The fractions of thylakoids isolated from intact plastids consisted of relaxed thylakoid lamellae. No other cellular membranes were recognized on transmission electron micrographs (Fig. 2D). Multiple (at least two) membrane-containing vesicles were observed in the envelope fractions (Fig. 2, E and F), which is in agreement with previous studies, in which the exposure of plastids to osmotic shock resulted in disruptions of their envelopes and their subsequent fusion (Douce and Joyard, 1980; Block et al., 1983). The tiny granules in Figure 2F may represent plastidial ribosomes.
Transmission electron micrographs of organellar and subplastidal fractions. A, Mitochondrial fraction isolated on a Suc gradient. E. gracilis mitochondria without other organellar contamination is seen. Precipitates of uranyl acetate used for membrane contrasting are visible between mitochondria, and poorer preservation of mitochondrial cristae is an artifact of the sample preparation. B, Plastid fraction isolated on a Percoll gradient containing intact plastids (P), holes in the resin after paramylon grains (PG), and a small amount of pellicle residue (PR). C, Two electron micrographs of isolated plastids (P), each containing three envelope membranes shown by red arrows. D, Relaxed thylakoid lamellae in the thylakoid fraction obtained from osmotically shocked intact plastids. E, Plastid envelope fraction obtained from osmotically shocked intact plastids consisting of tiny vesicles of various size and plastid ribosomes (black dots). F, Enlarged section of E.
Two different chemical analytical approaches were employed to exclude contamination in the fractions. Since it is well known that the occurrence of chlorophylls in the cell is limited to the chloroplast and, on the other hand, cardiolipins (CLs) are present only in mitochondria, we chose these molecules as markers of contamination. Chlorophyll a and chlorophyll b were found only in plastid fractions and thylakoid subfractions. The ratio of both chlorophylls is comparable between the two fractions. No chlorophyll was detected in envelope subfractions or mitochondrial fractions (Supplemental Fig. S2). A high content of pheophytin (a chlorophyll intermediate) was found accompanying chlorophylls. Tetrapyrrole compounds were measured by HPLC HR-MS/MS. The determination of CL content was more complicated, because CLs possess very low ionization performance in the positive mode (Hsu et al., 2005). Therefore, CLs were separated from the other compounds present by TLC and detected spectrophotometrically after derivatization. The TLC technique revealed CLs only in the mitochondrial fraction, at 12.4% ± 1.9% of the level of detected structural lipids (Supplemental Fig. S3). No CLs were detected in other cellular fractions. Although we were not able to determine the CL content in the E. gracilis envelopes due to limited sample amounts, we can indirectly rule out the possibility of contamination by mitochondria, because CLs were missing in the whole-plastid fraction, from which envelopes were separated. In summary, both TEM and chemical analysis suggest that plastid fractions were not contaminated by whole mitochondria and mitochondrial fractions were not contaminated by whole plastids or thylakoids. Strictly speaking, though, we cannot exclude contamination of mitochondria by plastid envelopes and contamination of plastids by outer mitochondrial membranes, as we do not have specific markers for these membrane types.
Mitochondria of E. gracilis Consist of PLs and CLs
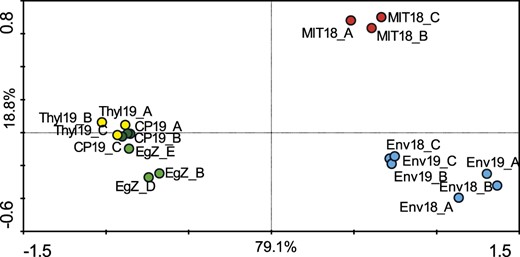
There were apparent differences in the composition of structural lipids between mitochondria, plastids, and plastid membranous subfractions (Fig. 3). The data from whole-cell, plastid, and thylakoid isolates clustered together, demonstrating that thylakoids were the dominant membranes in the E. gracilis cells, while mitochondrial and plastid envelope fractions formed two separate clusters, showing significant differences among these membrane types (Monte Carlo test, P = 0.001). A deeper look, focused on detailed lipid composition, explained the segregation of data (Supplemental Fig. S4).
A principal component analysis (PCA) ordination biplot (first and second ordination axes) showing similarity among the lipid record (HPLC HR-MS/MS) of E. gracilis strain Z and its organellar (sub)fractions. The distribution of data explains variability in the data set by 79.1% on the x axis and 18.8% on the y axis (meaning that differences among samples are more substantial in horizontal distances than in vertical distances). Redundancy analysis (RDA) and Monte-Carlo permutation test (unrestricted permutations; n = 999) confirmed significant differences among the obtained data sets (P = 0.001). The ordination plot shows that the thylakoid subfraction (yellow) is much more similar to whole cells (light green) and to the plastid fraction (dark green) but is quite different from the mitochondrial fraction (red) or the envelope subfraction (blue). Since thylakoid membranes are highly abundant in photosynthetic plastids, the data distribution is expected and also shows the accuracy of the methodology used for membrane fractionation. CP, Plastid fractions; EgZ, E. gracilis strain Z; Env, plastid envelope subfractions; MIT, mitochondrial fractions; Thyl, thylakoid subfractions.
The predominant mitochondrial lipids are PLs (Fig. 4A), with PC being the most abundant lipid class, followed by PE (Fig. 4B) and then by the hallmark mitochondrial lipid CL (12.4% ± 1.9%; Supplemental Fig. S3). A higher amount of PE-Os was recorded in mitochondrial fractions compared with plastid ones. We also detected considerable amounts of MGDGs in the mitochondrial fractions (Fig. 4B). The majority of PLs in mitochondria consisted of short saturated or monounsaturated FAs (C14:0, C16:0, and C16:1) in the sn-2 position accompanied by longer and unsaturated FAs (C20:3, C20:4, and C20:5) in the sn-1 position (Supplemental Fig. S5; Supplemental Table S3). This architecture is reflected by the ratio of FA species obtained by GC-FID (Fig. 4C).
Lipid and FA composition of E. gracilis cells (EgZ), plastids (CP), and mitochondria (Mit). The data were obtained by HPLC HR-MS/MS, and three biological replicates were measured for each E. gracilis cellular fraction. sd values are expressed as error bars. A, Bar graph showing percentages of coupled lipid classes. GDAGs include DGDGs, MGDGs, and SQDG; PLs include PCs, PE-Os, PEs, and PIs. B, Bar graph showing percentages of detected lipid classes. Data were obtained by coupling peak areas of intact lipid species. C, Bar graph showing percentages of selected fatty acid methyl esters (FAMEs) detected by GC-FID analyses (n = 3). Depicted FAs represent approximately 80% of all detected FAMEs in all cellular fractions.
The Lipid Composition of E. gracilis Plastids and Thylakoids Is Similar and Differs from That of Plastid Envelopes
E. gracilis plastids contain mostly GDAG lipids, and their lipid profile is comparable to the lipid composition of whole E. gracilis cells (Figs. 3 and 4, A and B). Although the distribution of GDAG classes in plastids favored MGDGs (Fig. 4B; Supplemental Fig. S5), the overall representation of GDAGs in plastids and in whole cells was similar (Fig. 4A). Also, the level of PE-Os was lower in plastid fractions; on the other hand, the isolated plastids were enriched in PCs compared with whole cells (Fig. 4B), which might be explained by an increased PC content in plastid envelopes (Fig. 5B). As mentioned above, the dominant FAs in plastid fractions were short PUFAs (C16:3n3 and C18:3n3; Fig. 4C). Very interesting is the substantial increase of MGDG 18:3/16:3, from 1.34% ± 0.5% in whole cells to 18.83% ± 1.2% in plastid fractions (Supplemental Fig. S5). Additionally, an obvious difference in the content of PE O-16:0/18:1 was recorded between mitochondria and plastids (Supplemental Fig. S5).
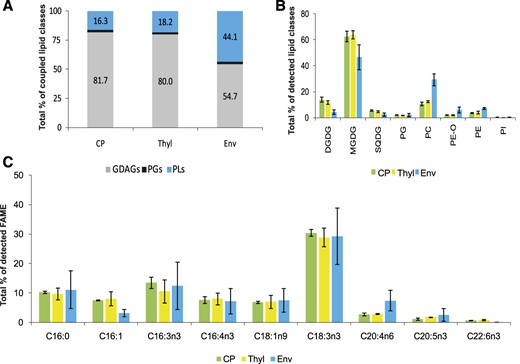
Lipid and FA composition of E. gracilis plastids (CP), thylakoids (Thyl), and plastid envelopes (Env). The data were obtained by HPLC HR-MS/MS, and three biological replicates were measured for each E. gracilis (sub)plastidial fraction. sd values are expressed as error bars. A, Bar graph showing percentages of coupled lipid classes. Data were obtained by coupling peak areas of intact lipid species. GDAGs include DGDGs, MGDGs, and SQDG; PLs include PCs, PE-Os, PEs, and PIs. B, Bar graph showing percentages of detected lipid classes. Data were obtained by coupling peak areas of intact lipid species. C, Bar graph showing percentages of selected fatty acid methyl esters (FAMEs) detected by GC-FID analyses (n = 3). Depicted FAs represent approximately 80% of all detected FAMEs in all (sub)plastidial fractions.
The multivariate statistical analysis clearly showed that the overall lipidomic profile in thylakoid fractions was similar to that in whole plastids (Figs. 3 and 5), underlining a situation in photosynthetically active plastids in which thylakoids form the bulk of the plastid membranes. Increased diversity in the composition of lipid species is remarkable in the envelope fractions (Fig. 3), where GDAGs represented only about half of all structural lipids (Fig. 5, A and B). The most enriched lipid classes in the plastid envelopes compared with thylakoid fractions were PCs and PE-Os (Fig. 5B). Surprisingly, we detected the enrichment of PEs in E. gracilis plastid envelopes (Fig. 5B). Also, the profile of FAs acquired by GC-FID shows a high level of similarity in all three plastid subfractions. The predominant FAs were 18:3n3, followed by 16:3n3 and 16:0. The increased proportion of C20:4 underlines the increasing number of PCs in the envelope fractions (Fig. 5C; Supplemental Table S3).
DISCUSSION
Lipid and FA Profiles of E. gracilis Cells and the Presence of GDAGs in W10 and OFL Mutants
The lipid profiles of E. gracilis whole cells were similar to those reported by Shibata et al. (2018). However, the distribution of structural lipids was slightly different: the ratio of MGDGs and DGDGs, almost 1:1 in our measurements (Fig. 1C), showed a higher proportion of MGDGs in the earlier study (Shibata et al., 2018). High levels of DGDGs in the cells were reported in plants during phosphorus deprivation. Under this condition, DGDGs are incorporated into mitochondrial membranes to supplement the lack of PLs (Jouhet et al., 2004; for review, see Block et al., 2007). However, we do not expect this situation in our cells because, similar to Shibata et al. (2018), we cultivated E. gracilis cells in modified Cramer-Myers medium (Cramer and Myers, 1952) containing phosphates. Moreover, DGDGs are almost absent from the mitochondrial fraction (Fig. 4B). Hence, we ascribe the change to different light conditions during cultivation.
The occurrence of PE-O lipid species in a mixotrophic E. gracilis is surprising. In animals, PE-Os are precursors for 1-alk-1-enyl-2-acyl-sn-glycerol-3-PEs, trivially called plasmalogens (Garg and Haerdi, 1993) and platelet-activating factors (Wanders and Brites, 2010). However, no plasmalogens were detected in our lipid extracts, corroborating the pioneering study focusing on plasmalogens in etiolated E. gracilis (Helmy et al., 1967). Previously, PE-Os were found in heterotrophic euglenozoans, such as Trypanosoma cruzi and Leismania donovani; however, phototrophs were hypothesized to lack these lipids (Thompson and Nozawa, 1972). Contrarily, Meneses et al. (1993) detected plasmalogens in the marine algae Bryothamnion triquetrum and Padyna gymnospora using NMR. Since PE-Os are only known as plasmalogen precursors (Dean and Lodhi, 2018), the presence of PE-Os in the above-mentioned organisms is expected. Ether lipids play a role in physiochemical properties of membranes (favoring membrane fusion), as receptor mediators, and as antioxidants (Garg and Haerdi, 1993; Horvath and Daum, 2013). Identification of PE-Os in E. gracilis was achieved by high-resolution data (Supplemental Table S1) and by the multiple ion fragmentation on third and fourth level (MS3 and MS4), the same fragmentation pattern was described by Hsu and Turk (2007; for detailed information, see Supplemental Fig. S6) and compared with proposed fragments in the LIPID MAPS structure database. A greater proportion of this lipid in bleached mutants (Fig. 1C) and in the mitochondrial fractions (Fig. 4B) supports the hypothesis that PE-Os are components of mitochondrial membranes and other cell membranes outside plastids. For instance, in the endoplasmic reticulum, known for rich membrane traffic, PE-Os could play a role in membrane fusion events. Furthermore, ether lipid synthesis finishes in the endoplasmic reticulum (Garg and Haerdi, 1993; Munn et al., 2003; Dean and Lodhi, 2018). Indeed, PE-Os could also be components of the outer plastid envelope, reflecting heavy membrane traffic between endoplasmic reticulum and plastids.
Our study presents the overall intact lipid composition of mixotrophic E. gracilis cells. Previous research focused only on intact DGDGs and MGDGs (Craig et al., 2015) identified the same set of GDAG species, but with different ratios. It has been proven many times by many authors that FA composition is dependent on multiple factors, such as growth phase (Schwarzhans et al., 2015), temperature (Craig et al., 2015), light intensity (Constantopoulos and Bloch, 1967), and carbon-nitrogen balance (Regnault et al., 1995).
Previous studies dealing with the FA composition of autotrophic E. gracilis cultures identified a little more than 20 FAs in total, ranging from lauric acid (C12:0) to lignoceric acid (C24:0; Hulanicka et al., 1964; Constantopoulos and Bloch, 1967; Regnault et al., 1995; Shibata et al., 2018). In this study, we detected 22 FA species, which include all those previously reported. A high amount of short PUFAs, such as C16:3, C16:4, and C18:3, has been recorded in all previously examined autotrophic or mixotrophic E. gracilis cultures, reflecting their presence in GDAGs (Shibata et al., 2018), which we also support. The presence of C20:5 and C22:6 FAs has been reported by most authors, except for Rosenberg and Pecker (1964) and Constantopoulos and Bloch (1967).
The OFL and W10 bleached mutants of E. gracilis were for a long time considered to be organisms with a laboratory-induced absence of plastids, supported by the absence of plastid structures (Osafune and Schiff, 1983; Polónyi et al., 1998). Furthermore, W10 was shown to lack a plastid-specific sulfolipid, SQDG (Saidha and Schiff, 1989). We have confirmed the absence of SQDG in W10 cells; however, we identified small amounts of other GDAGs in this strain, mainly MGDGs (Fig. 1C). All three GDAGs (including SQDG) were detected in the strain OFL, dominated by DGDG (Fig. 1, B and C). As plastids are the only known places of GDAG synthesis and are their major repositories, the presence of GDAGs in both bleached mutants indicates that at least some plastid remnants persist in their cells. For instance, Euglena longa possesses a nonphotosynthetic plastid that is undetectable by microscopic techniques, even by TEM (Kivic and Vesk, 1974). However, molecular biology has provided clear evidence for its presence (Gockel and Hachtel, 2000) as well as for the presence of a small amount of GDAGs (Füssy et al., 2020). On the other hand, there are no published data about the presence of plastid DNA in OFL and W10 mutants. The increase of PLs in relation to GDAGs in both bleached strains resembles the situation in strain Z grown in the dark (Rosenberg et al., 1966; Matson et al., 1970; Shibata et al., 2018).
Lipid and FA Profiles of E. gracilis Mitochondria and Plastids
We determined PCs and PEs to be the major structural lipids of isolated E. gracilis mitochondria, which is consistent with the situation in model species mitochondria (Horvath and Daum, 2013) and in agreement with the results of Shibata et al. (2018), who detected PCs and PEs as the most abundant lipids in a dark-grown E. gracilis culture with well-developed mitochondria. It is also reflected in the FA composition of mitochondria (Fig. 4C). C16:0, C20:4, and C20:5 are the most abundant FAs originating from the PLs; on the other hand, short PUFAs such as C16:3 and C18:3 originating from GDAGs and PGs are less abundant (Supplemental Table S2). Interestingly, plasmalogens of these lipid classes have been reported in mitochondria of specialized animal tissues, such as the heart or brain (Horvath and Daum, 2013). This finding supports our hypothesis that PE-Os are enriched in the mitochondrial membranes (Fig. 4B). Furthermore, we detected remarkable amounts of MGDGs (Fig. 4B) in mitochondrial fractions. Rather than real constituents of mitochondrial membranes, we consider these to be contamination from the plastid fraction. Supplemental Tables S2 and S3 and Supplemental Figure S4 show detailed lipid and FA composition of studied E. gracilis strains and cellular (sub)fractions. The most abundant MGDGs detected in mitochondria contain 20:3/16:4, 20:5/16:1, and 20:4/16:1 FA chains. If these lipids were real components of mitochondrial membranes, we would expect them to be abundant in the E. gracilis mutants also. However, this is not the case, because the detected GDAGs in the OFL mutant are DGDGs with 18:(1-3)/16:(1-4), MGDG 16:1/16:1, and SQDGs with 16:0/16:(0-1) and 14:0 FA chains; and in the W10 mutant, DGDGs with 18:3/16:(2-3) and MGDGs with 16:1/16:1 and 18:(1-2)/16:1 FA chains were detected the most. These lipids are typical components of plastid membranes in general and were detected in E. gracilis plastids too. Furthermore, MGDGs have never been reported as components of mitochondrial membranes, unlike the DGDGs that accumulate in plant mitochondria under phosphorus-deficient conditions. As we did not have any visible proof of plastid contamination on electron micrographs (Fig. 2A) and we did not detect any chlorophyll in the mitochondria (Supplemental Fig. S2), we assume that the MGDGs found in the mitochondria belong to plastid envelope remnants that stuck onto mitochondrial membranes after cell disruption during the isolation procedure.
The main differences in the lipid composition of E. gracilis plastids and whole cells are varying ratios of MGDGs and DGDGs (Fig. 4B). It has been shown that DGDGs play a crucial role in the formation of thylakoid membranes. The MGDG-to-DGDG ratio influences the transition from hexagonal to lamellar organization, allowing plants to quickly respond to changing conditions (Demé et al., 2014). We suppose that stress caused to plastids during isolation could have resulted in the changes in MGDG-to-DGDG ratios between isolated plastids and whole E. gracilis cells. The relevance of plastid isolation is underlined by the FA profiles of whole cells and plastid fractions. The increasing ratio of short PUFAs (C16:3, C16:4, and C18:3) at the expense of C16:0 and C18:1 (Fig. 4C) in isolated plastids follows the results of Shibata et al. (2018), where short PUFAs were determined to be the main FAs in E. gracilis GDAGs.
Lipid Composition of E. gracilis Thylakoids and Plastid Envelopes
HPLC HR-MS/MS and PCA showed that the E. gracilis thylakoid lipid profile pattern follows the lipid profile of whole plastids, which corresponds to the fact that thylakoids form the largest part of plastid membranes (Figs. 3 and 5). Surprisingly, we identified two classes of lipid compounds, PE-Os and PEs, in whole plastids and in the plastid subfractions (in both thylakoids and envelopes; Figs. 4B and 5B) that have never been reported in primary plastids. This fact may result from cross-contamination with other cellular membranes (Mendiola-Morgenthaler et al., 1985; Block et al., 2007), although we can exclude contamination by the inner mitochondrial membrane due to the lack of CLs in plastid fractions (Supplemental Fig. S3). Indeed, pellicle residue was seen in transmission electron micrographs of isolated plastids (Fig. 2B), and the most abundant lipid class of the E. gracilis pellicle was determined to be PE (Nakano et al., 1987). The presence of the PE-Os could be explained by heavy membrane traffic between the plastid and endoplasmic reticulum. Therefore, this may reflect a genuine difference from primary plastids caused by the presence of the additional membrane derived from the eukaryotic endomembrane system, the outermost envelope.
Can We Infer the Origin of the Middle Plastid Envelope Based on Its Structural Lipid Composition?
Unlike any other cell membranes, those of primary plastids are known to have a very specific composition of structural lipids, dominated by GDAGs. Should the middle plastid envelope be derived from the primary plastid, it should be expected to be rich in GDAGs too. As the innermost envelope tends to have the same lipid composition as thylakoids (figure 1A in LaBrant et al., 2018) and the percentage of GDAGs in envelopes drops by one-third from the thylakoid value, from 80% to 54.7% (Fig. 5A), this suggests that one (presumably the outermost) membrane is GDAGs free and contains PLs while the remaining two are enriched by GDAGs. Put another way, if GDAGs were to be present only in the innermost envelope membrane, their fraction in the envelopes could reach a maximum of 33% if this membrane’s lipids were 100% composed of GDAGs. The 54.7% content determined by us therefore indicates that GDAGs are present in more membranes, most likely in the two innermost. This, of course, assumes that the ratio of the three membranes in our fraction is 1:1:1, which may not be true. Contamination by thylakoids can be excluded, because we did not detect any chlorophyll in the envelope fraction. On the other hand, we cannot exclude the possibility that a portion of the purified plastids lost their outermost membrane(s) during preparation, which would artificially increase the GDAG composition. Indeed, in the extreme case, in which the outermost membranes were completely stripped off and the innermost membranes contained 100% GDAGs, we could observe a 54.7% GDAG content in the fraction even if the middle membranes were free of GDAGs. This is, however, a very unlikely situation, and we are able to distinguish all three membranes in at least some plastids by TEM (Fig. 2C). Even with these unknown factors in mind, we conclude that our observation is best explained by the hypothesis that the middle membrane contains GDAGs. The simplest explanation for GDAG presence would be an origin from the primary plastid. Still, the lipid content of the membrane may also follow functional constraints, and for this reason, we cannot exclude the scenario in which the middle membrane, although originating from the cytoplasmic membrane of the symbiont, was secondarily enriched in GDAGs. Finally, a plausible way by which to decrease the number of membranes is the fusion of the two neighboring middle (second and third) membranes, the result of which would then have a mixed origin. Whatever is the evolutionary implication, our results suggest that, in terms of lipid content, the middle plastid envelope membrane of E. gracilis resembles more closely the membranes of the primary plastid than other eukaryotic membranes. Applying this concept to other secondary plastid-bearing organisms and the lipid composition of their plastids and envelopes would be very interesting.
MATERIALS AND METHODS
Cultivation
Cultures of the pigmented Euglena gracilis strain Z and the bleached mutants W10 and OFL were cultivated in modified Cramer-Myers medium (Cramer and Myers, 1952) supplemented with 0.4% (v/v) ethanol and pH adjusted to 6.9. Before inoculation, vitamin B1 and vitamin B12 were aseptically added into the medium at final concentrations of 0.1 μg mL−1 and 0.5 ng mL−1, respectively. Cultures were synchronously grown (light/dark cycle = 12/12 h) for 1 week, were harvested 2 h after entering the light stage, and were used for lipid analysis of whole cells and for isolation of mitochondria (from strain Z only).
Cultures of E. gracilis strain Z used for the isolation of whole plastids and thylakoid membrane and plastid envelope membrane subfractions were cultivated synchronously for 1 week in 1 to 2 L of the modified Cramer-Myers medium (Cramer and Myers, 1952) under the same conditions as previously, except that the concentration of vitamin B12 in the medium was decreased to 0.05 ng mL−1 to ensure high plastid number and low paramylon content (Vacula et al., 2007).
Isolation of Mitochondria, Plastids, and Plastid Subfractions of E. gracilis
Isolation of mitochondria was performed according to the protocol described by Dobáková et al. (2015). The mitochondrial fraction was snap frozen in liquid nitrogen for lipid analysis or fixed for TEM.
A protocol for plastid isolation was established by combining two published protocols (Napier and Barnes, 1995; Vacula et al., 2007). Briefly, cells were harvested and washed two times in 1× GRM buffer (330 mm sorbitol, 50 mm HEPES-KOH, pH 8, 2 mm EDTA, 1 mm MgCl2, and 1 mm MnCl2; Napier and Barnes, 1995) without protease inhibitors. The cells were resuspended in 1× GRM buffer with plant protease inhibitor cocktail (PIC; Sigma-Aldrich) and disrupted using a French press at 1,500 psi (Vacula et al., 2007). The crude cell lysate was diluted with 1× GRM + PIC and centrifuged for 3 min at 250g. The supernatant was transferred into a sterile tube, and the centrifugation step was repeated several times to remove unbroken cells and debris. The supernatant was then centrifuged at 2,600g for 5 min, and the pellet, containing plastids, was loaded on a Percoll (Sigma-Aldrich) gradient and centrifuged at 5,000g in a swinging-bucket rotor (Beckman Coulter SW-41Ti) for 20 min at 4°C (Napier and Barnes, 1995). The bottom green layer, containing intact plastids, was removed from the gradient, diluted with 1× GRM without PIC, and centrifuged at 2,600g for 5 min. All steps were performed on ice or in precooled centrifuges. The collected plastids were washed, and the plastid pellet was snap frozen in liquid nitrogen for subsequent lipid analysis, fixed for TEM, or used further for the thylakoid and envelope isolations.
Thylakoids and plastid envelopes were isolated according to Peltier et al. (2000) from the intact plastids by exposure to osmotic shock by resuspending the plastids in 50 mm Tris-HCl and 5 mm MgCl2, pH 8, while resting on ice for 10 min. The sample was then centrifuged at 4°C and 10,000g for 10 min. The supernatant, containing plastid envelopes, was kept on ice, and the thylakoid pellet was washed twice in 10 mm Tris-HCl, pH 8, and centrifuged. The supernatant containing envelope membranes was finally centrifuged on the maximum g in a table centrifuge at 4°C for 40 min. The plastid subfractions were frozen in liquid nitrogen for lipid analysis or fixed for TEM.
TEM
(Sub)organellar fractions were fixed in 2.5% (v/v) glutaraldehyde (Electron Microscopy Science [EMS]) buffered with 0.1 m sodium cacodylate, pH 7.4 (Sigma-Aldrich), for 1 h at room temperature. After washing in 0.1 m sodium cacodylate buffer, the fractions were postfixed and contrasted using 1% (v/v) osmium tetroxide (EMS) for 1 h on ice. Dehydration in a graded ethanol series (Penta Chemicals), each for 10 min on ice, and finally in absolute ethanol for 15 min, was followed by Epon resin (EmBed 812, medium grade; EMS) infiltration at 1:1 and 1:2 ratio with ethanol for 30 min each. Complete Epon resin infiltration was performed overnight with two other Epon changes, and resin polymerization occurred at 60°C for 48 h. Ultrathin sections (70 nm) were prepared on a Leica EM UC7 ultramicrotome (Leica Microsystems) and collected onto carbon- and Formvar-coated copper grids. The sections were postcontrasted with 4% (w/v) aqueous uranyl acetate solution (EMS) for 1 h at room temperature and 2% (w/v) lead citrate (EMS) for 10 min on ice and imaged at 80 kV using a JEOL 1011 TEM device equipped with a Veleta CCD camera and Olympus software (Soft Imaging Solutions).
Extraction of the Sample
Lipids were extracted from whole cells of E. gracilis strains and from subcellular fractions of strain Z with a chloroform and methanol solution (ratio 2:1) following the method of Folch et al. (1957). Samples were homogenized in the extraction solution with glass beads using a TissueLyser LT mill (Qiagen). Homogenates were dried and stored at −20°C for further analyses.
Intact Lipid Analysis by HPLC Electrospray Ionization-MS/MS and HPLC Electrospray Ionization-HR-MS/MS
Dried extracts were resolved in 500 µL of chloroform and methanol (1:2). One hundred microliters of the solution was mixed with an internal standard, PC 17:0/17:0 (20 µL mL−1; Sigma-Aldrich). The sample aliquots (5 µL) were injected by an Accela autosampler (Thermo Fisher Scientific) and separated on a Gemini column measuring 250 × 2 mm, internal diameter 3 µm (Phenomenex). A linear ion trap LTQ-XL mass spectrometer (Thermo Fisher Scientific) was used in both positive and negative ion electrospray ionization modes. The settings of the system followed methodology published earlier (Tomčala et al., 2017). Particular lipid species were determined based on mass-to-charge ratio value, retention time, behavior in positive and negative ionization modes, characteristic fragmentation pattern, and high-resolution data obtained by the mass spectrometry system, powered by an Orbitrap Q-Exactive Plus with a Dionex Ultimate 3000 XRS pump and a Dionex Ultimate 3000 XRS Open autosampler (all by Thermo Fisher Scientific) following the settings described by Tomčala et al. (2017). The identification of a particular lipid class as plasmenyl or plasmanyl was achieved by MS3 and MS4 experiments, in which collision energy was set to 35%; isolation width was set to 4 D in both mass spectrometry levels. The data from both spectrometers were acquired and processed using Xcalibur software version 2.1 (Thermo Fisher Scientific; for details see Tomčala et al., 2017).
TLC
TLC was used to investigate lipid class composition. The analysis was performed according to the method of Zahradníčková et al. (2014) with minor changes. TLC plates (20 × 10 cm, Silica Gel 60, 0.2-mm layer; Merck) were employed as a stationary phase. The plates were prewashed consecutively with chloroform:methanol (2:1) and with hexane. The lipid extracts were rediluted in 400 µL of methanol. Nitrogen was used as a spray gas, and 7 μL of the sample was applied with a CAMAG TLC Sampler 4 (Camag). The lipid classes were separated on TLC plates in a Twin Through Chamber 20×20 (Camag) using 10 mL of hexane:diethyl ether:acetic acid (80:18:2) as the mobile phase three times to remove nonpolar lipids, such as triacylglycerols and diacylglycerols, for the length of 8.5 cm. First, elution by chloroform:methanol:acetic acid:water (50:30:4:6) for 4.5 cm was used to separate the PLs and GDAGs. Then, chloroform:methanol:acetic acid:water (40:40:4:10) developed the TLC plate for 8 cm to segregate particular GDAG lipid classes. The chamber saturation was increased by filter paper in the development chamber (Camag). Plates were removed from the chambers and subsequently dried and sprayed with a solution of 3% (w/v) cupric acetate in 8% (w/v) phosphoric acid. The plate was then charred for 20 min at 160°C. Quantitative analysis of the separated lipid classes was performed by a CAMAG TLC Scanner 3 (Camag). The scanning was performed at a wavelength of 350 nm. Identification of the lipid classes was performed by comparison with an external standard (MGDG, DGDG, and SQDG [Larodan]; PG, PC, PE, and CL [Sigma-Aldrich]).
FA Analysis by GC-FID
Total lipid extracts for FA analyses were methylated according to the methods of Appelqvist (1968). FA composition was analyzed by GC (Trace Ultra FID; Thermo Scientific) using a BPX-70 50-m fused silica capillary column (internal diameter 0.22 mm, 0.25 μm film thickness; SGE). The initial temperature was 70°C and was held for a half minute, after which the temperature was raised by 30°C min−1 until reaching 150°C. After that, the temperature rose to 220°C by a rate of 1.5°C min−1 and was then held for 11 min. The whole analysis took approximately 60 min. The peaks were identified by comparing sample retention times with retention times of the standard mixture Supleco 37 Component FAME Mix (Sigma-Aldrich).
Multivariate Statistical Analysis
The data obtained from the areas of intact lipids or particular FA peak areas were statistically evaluated using ordination methods as follows: for all data, detrended correspondence analysis; for linear data, PCA, RDA, and Monte-Carlo permutation test (unrestricted permutations; n = 999); and for unimodal data, CA, CCA, and Monte-Carlo permutation test (unrestricted permutations; n = 999). The data were transformed by using the internal standard peak area of the particular sample followed by recalculation to the ratio of particular detected lipids. For each, the transformed peak areas were calculated in the deconvoluted total cell and peak areas. In the canonical analyses (RDA and PCA), the E. gracilis strains or cell fractions stood as categorical predictors. The Monte-Carlo permutation test was used for statistical significance determination. Statistics software CANOCO 4.5 (Biometrics, Plant Research International) was used for the detrended correspondence analysis, CA, CCA, PCA, RDA, and Monte-Carlo permutation test analyses.
Supplemental Data
The following supplemental materials are available.
Supplemental Figure S1. CA of obtained lipidomic records from different E. gracilis strains.
Supplemental Figure S2. Comparison of abundances of chlorophyll a and b and pheophytin, the chlorophyll precursor and also degradation product in E. gracilis strain Z cellular (sub)fractions.
Supplemental Figure S3. Comparison of abundances of cardiolipin in E. gracilis plastid, thylakoid, and mitochondrial fractions.
Supplemental Figure S4. Lipid and FA composition of E. gracilis strain Z, plastid envelopes, and mitochondria.
Supplemental Figure S5. Graph of representative intact lipid species in E. gracilis cells, plastids, and mitochondria.
Supplemental Figure S6. Fragmentation pattern of compound mass-to-charge ratio = 702 obtained from lipidic extract of E. gracilis: MS2, MS3, and MS4 experiments.
Supplemental Table S1. List of detected structural lipids in mixotrophic E. gracilis strain Z obtained by HPLC HR-MS/MS.
Supplemental Table S2. List of detected structural lipids in mixotrophic E. gracilis strain Z and two bleached mutants, OFL and W10.
Supplemental Table S3. List of intact structural lipid species in E. gracilis strain Z plastid, thylakoid, envelope, and mitochondrial fractions.
ACKNOWLEDGMENTS
We acknowledge the Imaging Methods Core Facility at BIOCEV for its support with obtaining the imaging data presented in this article.
LITERATURE CITED
Author notes
This work was supported by the Czech Science Foundation (grant no. 16–25280S) and by the Ministry of Education, Youth, and Sports of the Czech Republic (grant nos. LQ1604, CZ.1.05/1.1.00/02.0109, LM2018099, CZ.1.05/2.1.00/19.0380, CZ.02.1.01/0.0/0.0/16_025/0007370, and CZ.02.1.01/0.0/0.0/16_019/0000759).
Author for contact: vlada@natur.cuni.cz.
Senior author.
The author responsible for distribution of materials integral to the findings presented in this article in accordance with the policy described in the Instructions for Authors (www.plantphysiol.org) is: Vladimír Hampl (vlada@natur.cuni.cz).
V.H. and L.T. designed the experiments; L.T. performed cellular and organellar fractionation; A.T. carried out lipid analyses; all authors analyzed and interpreted data; L.T. and A.T. wrote the article; V.H. and M.O. supervised and complemented the article.