Abstract
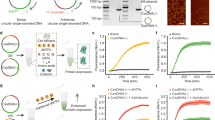
Improving the carboxylation properties of Rubisco has primarily arisen from unforeseen amino acid substitutions remote from the catalytic site. The unpredictability has frustrated rational design efforts to enhance plant Rubisco towards the prized growth-enhancing carboxylation properties of red algae Griffithsia monilis GmRubisco. To address this, we determined the crystal structure of GmRubisco to 1.7 Å. Three structurally divergent domains were identified relative to the red-type bacterial Rhodobacter sphaeroides RsRubisco that, unlike GmRubisco, are expressed in Escherichia coli and plants. Kinetic comparison of 11 RsRubisco chimaeras revealed that incorporating C329A and A332V substitutions from GmRubisco Loop 6 (corresponding to plant residues 328 and 331) into RsRubisco increased the carboxylation rate (kcatc) by 60%, the carboxylation efficiency in air by 22% and the CO2/O2 specificity (Sc/o) by 7%. Plastome transformation of this RsRubisco Loop 6 mutant into tobacco enhanced photosynthesis and growth up to twofold over tobacco producing wild-type RsRubisco. Our findings demonstrate the utility of RsRubisco for the identification and in planta testing of amino acid grafts from algal Rubisco that can enhance the enzyme’s carboxylase potential.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Atomic coordinates and structure factors for G. monilis Rubisco are accessible via Protein Data Bank (http://www.wwpdb.org) accession number 8BDB. All other data are available online as source files and via https://doi.org/10.6084/m9.figshare.21547515. Source data are provided with this paper.
References
Bailey-Serres, J., Parker, J. E., Ainsworth, E. A., Oldroyd, G. E. D. & Schroeder, J. I. Genetic strategies for improving crop yields. Nature 575, 109–118 (2019).
Simkin, A. J., López-Calcagno, P. E. & Raines, C. A. Feeding the world: improving photosynthetic efficiency for sustainable crop production. J. Exp. Bot. 70, 1119–1140 (2019).
Evans, J. R. & Lawson, T. From green to gold: agricultural revolution for food security. J. Exp. Bot. 71, 2211–2215 (2020).
Sharwood, R. E. Engineering chloroplasts to improve Rubisco catalysis: prospects for translating improvements into food and fiber crops. New Phytol. 213, 494–510 (2017).
von Caemmerer, S. Rubisco carboxylase/oxygenase: from the enzyme to the globe: a gas exchange perspective. J. Plant Physiol. 252, 153240 (2020).
Busch, F. A. Photorespiration in the context of Rubisco biochemistry, CO2 diffusion and metabolism. Plant J. 101, 919–939 (2020).
Evans, J. & Seemann, J. in Photosynthesis (ed. Briggs, W. R.) 183–205 (Alan R Liss, 1989).
Evans, J. R. & Clarke, V. C. The nitrogen cost of photosynthesis. J. Exp. Bot. https://doi.org/10.1093/jxb/ery366 (2018).
Iñiguez, C. et al. Evolutionary trends in RuBisCO kinetics and their co-evolution with CO2 concentrating mechanisms. Plant J. 101, 897–918 (2020).
Ermakova, M. et al. Installation of C4 photosynthetic pathway enzymes in rice using a single construct. Plant Biotech. J. 19, 575–588 (2021).
Long, B. M., Rae, B. D., Rolland, V., Förster, B. & Price, G. D. Cyanobacterial CO2-concentrating mechanism components: function and prospects for plant metabolic engineering. Curr. Opin. Plant Biol. 31, 1–8 (2016).
Fei, C., Wilson, A. T., Mangan, N. M., Wingreen, N. S. & Jonikas, M. C. Modelling the pyrenoid-based CO2-concentrating mechanism provides insights into its operating principles and a roadmap for its engineering into crops. Nat. Plants 8, 583–595 (2022).
Andersson, I. & Backlund, A. Structure and function of Rubisco. Plant Physiol. Biochem. 46, 275–291 (2008).
Iñiguez, C., Aguiló-Nicolau, P. & Galmés, J. Improving photosynthesis through the enhancement of Rubisco carboxylation capacity. Biochem. Soc. Trans. 49, 2007–2019 (2021).
Bracher, A., Whitney, S. M., Hartl, F. U. & Hayer-Hartl, M. Biogenesis and metabolic maintenance of Rubisco. Annu. Rev. Plant Biol. 68, 29–60 (2017).
Aigner, H. et al. Plant Rubisco assembly in E. coli with five chloroplast chaperones including BSD2. Science 358, 1272–1278 (2017).
Whitney, S. M., Baldet, P., Hudson, G. S. & Andrews, T. J. Form I Rubiscos from non-green algae are expressed abundantly but not assembled in tobacco chloroplasts. Plant J. 26, 535–547 (2001).
Zhu, X. G., Portis, A. R. & Long, S. P. Would transformation of C3 crop plants with foreign Rubisco increase productivity? A computational analysis extrapolating from kinetic properties to canopy photosynthesis. Plant Cell Env. 27, 155–165 (2004).
Lin, M. T. & Hanson, M. R. Red algal Rubisco fails to accumulate in transplastomic tobacco expressing Griffithsia monilis RbcL and RbcS genes. Plant Direct 2, e00045 (2018).
Gunn, L. H., Martin Avila, E., Birch, R. & Whitney, S. M. The dependency of red Rubisco on its cognate activase for enhancing plant photosynthesis and growth. Proc. Natl Acad. Sci. USA 117, 25890–25896 (2020).
Andersson, I. Catalysis and regulation in Rubisco. J. Exp. Bot. 59, 1555–1568 (2008).
Hanson, D. T. Breaking the rules of Rubisco catalysis. J. Exp. Bot. 67, 3180–3182 (2016).
Martin-Avila, E. et al. Modifying plant photosynthesis and growth via simultaneous chloroplast transformation of Rubisco large and small subunits. Plant Cell 32, 2898–2916 (2020).
Lin, M. T., Salihovic, H., Clark, F. K. & Hanson, M. R. Improving the efficiency of Rubisco by resurrecting its ancestors in the family Solanaceae. Sci. Adv. 8, abm6871 (2022).
Young, J. N. et al. Large variation in the Rubisco kinetics of diatoms reveals diversity among their carbon-concentrating mechanisms. J. Exp. Bot. 67, 3445–3456 (2016).
Whitney, S. M. & Sharwood, R. E. Construction of a tobacco master line to improve Rubisco engineering in chloroplasts. J. Exp. Bot. 59, 1909–1921 (2008).
Heureux, A. M. C. et al. The role of Rubisco kinetics and pyrenoid morphology in shaping the CCM of haptophyte microalgae. J. Exp. Bot. 68, 3959–3969 (2017).
Iñiguez, C., Galmés, J. & Gordillo, F. J. L. Rubisco carboxylation kinetics and inorganic carbon utilization in polar versus cold-temperate seaweeds. J. Exp. Bot. 70, 1283–1297 (2018).
Chen, Z. X., Yu, W. Z., Lee, J. H., Diao, R. & Spreitzer, R. J. Complementing amino acid substitutions within loop 6 of the alpha/beta-barrel active site influence the CO2/O2 specificity of chloroplast ribulose-1,5-bisphosphate carboxylase/oxygenase. Biochemistry 30, 8846–8850 (1991).
Bouvier, J. W. et al. Rubisco adaptation is more limited by phylogenetic constraint than by catalytic trade-off. Mol. Biol. Evol. 30, 2880–2896 (2021).
Wilson, R. H., Alonso, H. & Whitney, S. M. Evolving Methanococcoides burtonii archaeal Rubisco for improved photosynthesis and plant growth. Sci. Rep. 6, 22284 (2016).
Wilson, R. H., Martin-Avila, E., Conlan, C. & Whitney, S. M. An improved Escherichia coli screen for Rubisco identifies a protein–protein interface that can enhance CO2-fixation kinetics. J. Biol. Chem. 293, 18–27 (2018).
Zhou, Y. & Whitney, S. Directed evolution of an improved Rubisco; in vitro analyses to decipher fact from fiction. Int. J. Mol. Sci. 20, 5019 (2019).
Savir, Y., Noor, E., Milo, R. & Tlusty, T. Cross-species analysis traces adaptation of Rubisco toward optimality in a low-dimensional landscape. Proc. Natl Acad. Sci. USA 107, 3475–3480 (2010).
Tcherkez, G., Farquhar, G. & Andrews, T. Despite slow catalysis and confused substrate specificity, all ribulose bisphosphate carboxylases may be nearly perfectly optimized. Proc. Natl Acad. Sci. USA 103, 7246–7251 (2006).
Mueller-Cajar, O. et al. Structure and function of the AAA+ protein CbbX, a red-type Rubisco activase. Nature 479, 194–199 (2011).
Taylor, T. C. & Andersson, I. Structural transitions during activation and ligand binding in hexadecameric Rubisco inferred from the crystal structure of the activated unliganded spinach enzyme. Nat. Struct. Biol. 3, 95–101 (1996).
Chen, Z. X. & Spreitzer, R. J. Chloroplast intragenic suppression enhances the low CO2/O2 specificity of mutant ribulose-bisphosphate carboxylase/oxygenase. J. Biol. Chem. 264, 3051–3053 (1989).
Karkehabadi, S., Satagopan, S., Taylor, T. C., Spreitzer, R. J. & Andersson, I. Structural analysis of altered large-subunit loop-6/carboxy-terminus interactions that influence catalytic efficiency and CO2/O2 specificity of ribulose-1,5-bisphosphate carboxylase/oxygenase. Biochemistry 46, 11080–11089 (2007).
Gutteridge, S., Rhoades, D. F. & Herrmann, C. Site-specific mutations in a loop region of the C-terminal domain of the large subunit of ribulose bisphosphate carboxylase/oxygenase that influence substrate partitioning. J. Biol. Chem. 268, 7818–7824 (1993).
Parry, M. A. J., Madgwick, P., Parmar, S., Cornelius, M. J. & Keys, A. J. Mutations in loop six of the large subunit of ribulose-1,5-bisphosphate carboxylase affect substrate specificity. Planta 187, 109–112 (1992).
Galmés, J. et al. Environmentally driven evolution of Rubisco and improved photosynthesis and growth within the C3 genus Limonium (Plumbaginaceae). New Phytol. 203, 989–999 (2014).
Zhu, G., Jensen, R. G., Bohnert, H., Wildner, G. & Schlitter, J. Dependence of catalysis and CO2/O2 specificity of Rubisco on the carboxy-terminus of the large subunit at different temperatures. Photosynth. Res. 57, 71–79 (1998).
Joshi, J., Mueller-Cajar, O., Tsai, Y. C., Hartl, F. U. & Hayer-Hartl, M. Role of small subunit in mediating assembly of red-type Form I Rubisco. J. Biol. Chem. 290, 1066–1074 (2015).
Bhat, J. Y. et al. Mechanism of enzyme repair by the AAA+ chaperone Rubisco activase. Mol. Cell 67, 744–756.e6 (2017).
Genkov, T. & Spreitzer, R. J. Highly conserved small subunit residues influence Rubisco large subunit catalysis. J. Biol. Chem. 284, 30105–30112 (2009).
Spreitzer, R. J., Peddi, S. R. & Satagopan, S. Phylogenetic engineering at an interface between large and small subunits imparts land-plant kinetic properties to algal Rubisco. Proc. Natl Acad. Sci. USA 102, 17225–17230 (2005).
Whitney, S. M., Birch, R., Kelso, C., Beck, J. L. & Kapralov, M. V. Improving recombinant Rubisco biogenesis, plant photosynthesis and growth by coexpressing its ancillary RAF1 chaperone. Proc. Natl Acad. Sci. USA 112, 3564–3569 (2015).
Whitney, S. M. et al. Isoleucine 309 acts as a C4 catalytic switch that increases ribulose-1,5-bisphosphate carboxylase/oxygenase (Rubisco) carboxylation rate in Flaveria. Proc. Natl Acad. Sci. USA 108, 14688–14693 (2011).
Buck, S. et al. Escherichia coli expressing chloroplast chaperones as a proxy to test heterologous Rubisco production in leaves. J. Exp. Bot. 74, 664–676 (2023).
Sharwood, R. E., Ghannoum, O., Kapralov, M. V., Gunn, L. H. & Whitney, S. M. Temperature responses of Rubisco from Paniceae grasses provide opportunities for improving C3 photosynthesis. Nat. Plants 2, 16186 (2016).
Whitney, S. M. & Sharwood, R. E. Linked Rubisco subunits can assemble into functional oligomers without impeding catalytic performance. J. Biol. Chem. 282, 3809–3818 (2007).
Sharwood, R. E., Sonawane, B. V., Ghannoum, O. & Whitney, S. M. Improved analysis of C4 and C3 photosynthesis via refined in vitro assays of their carbon fixation biochemistry. J. Exp. Bot. 67, 3137–3148 (2016).
Whitney, S. M. & Sharwood, R. E. Rubisco engineering by plastid transformation and protocols for assessing expression. Methods Mol. Biol. 2317, 195–214 (2021).
Acknowledgements
This research was supported by the Australian Government through the Australian Research Council Centre of Excellence for Translational Photosynthesis CE140100015 and through the Formas Future Research Leaders grant 2017-00963. The work was also supported by the project ‘Structural dynamics of biomolecular systems (ELIBIO)’ (NO. CZ.02.1.01/0.0/0.0/15_003/0000447) from the European Regional Development Fund. We thank Diamond Light Source for access to beamline I02 (proposal MX11171-5) and B. Li for technical assistance.
Author information
Authors and Affiliations
Contributions
Y.Z., L.H.G. and S.M.W. conceived the project and all authors wrote the paper; L.H.G. and I.A. determined the crystal structure; L.H.G. and S.M.W. generated and analysed all enzymes; Y.Z. and R.B. generated the transgenic tobacco lines, performed the molecular and protein analyses, and measured photosynthesis and growth.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks Cong-Zhao Zhou and Thomas Sharkey for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Interactions of the GmRbcS βE-βF hairpin loop.
Interactions between residues within the βE-βF sheet (hairpin loop) of adjacent GmRbcS subunits (coloured teal, residue labeling in red and blue correspond to GmRbcS subunits B and C in PDB 8BDB, respectively) and residues in neighboring RbcL subunits C and A (coloured grey, labels in black). Residue contacts are indicated by dashed lines.
Extended Data Fig. 2 Quantifying Rubisco Ko.
(A) Linear response of Kc to increasing [O2] for E. coli produced RsRubisco, (B) RsRubisco-chi11 and (C) Rubisco from tobacco leaves. The Kc is derived from the y-axis (that is where [O2] = 0) and Ko calculated as Kc/slope of the linear fit. Fits were made to the mean Kc values (±S.E) derived from Michaelis-Menten fits to 14CO2 fixation assays measured using n = 3 or 4 independent biological samples per Rubisco isoform (each symbol representing a differing enzyme preparation). See Table 1 for Ko values.
Extended Data Fig. 3 Generation of the tobRr::X transforming master-line and tobRsL11S.
(A) Schematic summary of the Rubisco genetics and transformation process used by20 to generate the tobRsLS::X genotype from which a homozygous line coding a single cbbX allele was generated and used to pollinate the Rhodospirillum rubrum L2 RrRubisco tobRr genotype26 to produce (B) the heterozygous RsCbbX (RsRca) producing tobRr::X line with all the progeny (C) kanamycin resistant (see pBinTP-cbbX plasmid in panel A showing co-transformation of the nptII and cbbX alleles). (D) The RsRubisco bioengineering tobRr::X master-line was plastome transformed with pLEVRsL11S that contained plastome flanking sequence (see dashed lines with numbering relative to the corresponding tobacco plastome GenBank accession NC_001879.2 sequence) that directed the replacement of the rbcM gene with the Rs-rbcL-rbcS-aadA operon coding RsRubisco-chi11 that differs by 3 nucleotides (as shown) to the RsRbcL in the tobRsLS::X line produced by20. Shown are the location of the 5UTR-probe and the BglII sites and fragment size it hybridizes to. (E) 32P-5UTR-probe labelled genomic DNA-blot screen of 6 independent tobRsL11S::X lines following 2 rounds of spectinomycin selection, a tobRsLS::X line (positive control) and tobRr::X (negative control). Lines #2 and #5 were deemed homoplasmic as no rbcM containing plastome copies (5.5 kb fragment) were evident. (F) Subsequent native-PAGE screen of soluble protein from three tobacco (wt), three tobRsLS::X plants and three of the six T0 tobRsL11S::X lines grown to maturity for seed. The seed (T1 progeny) from tobRsL11S line #1::X was used in subsequent growth comparisons (Fig. 5).
Extended Data Fig. 4 Comparative Loop 6 interactions.
Comparative interactions around the GmRubisco Loop 6 site 1 (A331) and site 2 (V334) residues relative to (A, C) Rs and (B, D) SoRubisco. Residue contacts are indicated by dashed lines.
Supplementary information
Supplementary Information
Supplementary Protocols, Figs.1 and 2, Tables 1–3 and References.
Source data
Source Data Fig. 3
Unprocessed Coomassie-stained native PAGE.
Source Data for Figs. 3 and 5, and Extended Data Fig. 2
Spreadsheet-based data for Table 1, Figs. 3c and 5c–g, and Extended Data Fig. 2 combined into a workbook with multiple tabs.
Source Data Extended Fig. 3e
Unprocessed DNA blot (only low-resolution source image available) for Extended Data Fig. 3e and unprocessed Coomassie-stained native PAGE for Extended Data Fig. 3f.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhou, Y., Gunn, L.H., Birch, R. et al. Grafting Rhodobacter sphaeroides with red algae Rubisco to accelerate catalysis and plant growth. Nat. Plants 9, 978–986 (2023). https://doi.org/10.1038/s41477-023-01436-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-023-01436-7
This article is cited by
-
Understanding the biochemical, physiological, molecular, and synthetic biology approaches towards the development of C4 rice (Oryza sativa L.)
Cereal Research Communications (2024)
-
Theme and variations
Nature Plants (2023)