Abstract
While various QRS detection and classification methods were developed in the past, the Holter ECG data acquired during daily activities by wearable devices represent new challenges such as increased noise and artefacts due to patient movements. Here, we present a deep-learning model to detect and classify QRS complexes in single-lead Holter ECG. We introduce a novel approach, delivering QRS detection and classification in one inference step. We used a private dataset (12,111 Holter ECG recordings, length of 30 s) for training, validation, and testing the method. Twelve public databases were used to further test method performance. We built a software tool to rapidly annotate QRS complexes in a private dataset, and we annotated 619,681 QRS complexes. The standardised and down-sampled ECG signal forms a 30-s long input for the deep-learning model. The model consists of five ResNet blocks and a gated recurrent unit layer. The model's output is a 30-s long 4-channel probability vector (no-QRS, normal QRS, premature ventricular contraction, premature atrial contraction). Output probabilities are post-processed to receive predicted QRS annotation marks. For the QRS detection task, the proposed method achieved the F1 score of 0.99 on the private test set. An overall mean F1 cross-database score through twelve external public databases was 0.96 ± 0.06. In terms of QRS classification, the presented method showed micro and macro F1 scores of 0.96 and 0.74 on the private test set, respectively. Cross-database results using four external public datasets showed micro and macro F1 scores of 0.95 ± 0.03 and 0.73 ± 0.06, respectively. Presented results showed that QRS detection and classification could be reliably computed in one inference step. The cross-database tests showed higher overall QRS detection performance than any of compared methods.
Similar content being viewed by others
Introduction
The electrocardiograph (ECG) is a common method to analyse heart rhythm and its disturbances. While some arrhythmias (such as atrial fibrillation in Fig. 1C,D) may be episodical, a patient can be equipped with a Holter ECG device to record longer periods (from 24 h to several days or weeks). Alternatively, a patient may be equipped with a wearable device to check the rhythm permanently. In both scenarios, the entry point in clinical analysis of Holter ECG data is reliable beat (i.e., QRS complex) detection and classification (Fig. 1) to describe patient rhythm.
Examples of Holter ECG signals: Normal sinus rhythm (A); the same patient with the same rhythm, but disturbed by noise due to patient movements (B). A different patient with atrial fibrillation and premature ventricular contractions (PVC) in a couplet (C); the same patient a few minutes later, with disturbed ECG signal (D) with at least one PVC.
QRS detection
Existing beat detection methods are based on morphology analysis as QRS slope, amplitude, and width1, digital filtering2,3,4,5,6, wavelet transform7, machine learning8,9 or deep-learning10,11 approaches. These QRS detection methods performed great on public datasets as an MIT-BIH12. However, the Holter ECG data acquired during daily activities might still be challenging. The data contains a higher amount of noise caused by patient movements (Fig. 1B,D), further affected by the quality of electrode placement since during long-term Holter monitoring, subjects often place electrodes by themselves.
QRS classification
Knowing QRS positions allows for evaluating heart-rate variability, minima, maxima, or the presence of pauses. If more precise ECG analysis is required, the most common beat classes can be recognised (Fig. 2): normal beats, premature atrial contractions (PAC), or premature ventricular contractions (PVC). When these beat classes are known, pathologies formed by specific beat sequences can be analysed. Then, for example, we can automatically identify PVC couplets, triplets, sustained or persistent ventricular tachycardia, or runs of supraventricular tachycardia.
Existing classification approaches may be based on engineered features13,14 or, nowadays, on deep learning techniques15,16, further implementing convolutional17,18 or recurrent layers19,20. Most of them are trained and tested using the public MIT-BIH database (47 patients) or CPSC database (2000 patients), both recorded in resting supine position. These methods are supposed to work using preceding QRS detection. However, deep-learning techniques allow the preparation of models covering QRS detection and classification in one inference step. Thus, we present a robust, deep-learning method to detect and classify QRS complexes in ECG data recorded during usual daily activities. We also introduce a novel approach, delivering detection and classification results in one inference step.
Data
We used private (Fig. 3A) and public (Fig. 3B) ECG datasets in this study. The anonymised, private ECG dataset was collected during routine ECG screening and, therefore, was not subject to the ethical committee by Czech law. This private dataset was used for the method development (Fig. 3C) and testing, and public datasets were used only for cross-database tests (Fig. 3D). The lead "I" was used if the dataset contained multiple leads. If it was not present, the first ECG lead was used.
Dataflow in the presented study: private MDT data (A) were split into training, validation, and test subsets. Training and validation subsets were used to develop the proposed method (C). Next, the method performance was measured (D) using MDT private test data and data from twelve public databases (B). For the QRS classification task, only four databases were used.
Private dataset
The private dataset MDT (Medical Data Transfer, s. r. o., Brno, Czechia) consisted of 12,111 single-lead Holter ECG recordings. Each recording was 45 s long, sampled at 200 Hz. Recordings were acquired from patients during usual daily activities and often contained a high amount of noise (Fig. 1B,D). We have developed a software tool, "QRS Marker". Two specialists with more than five years of experience with QRS detection and classification semi-automatically marked 619,681 QRS complexes in this tool. Next, data were split into training (80%), validation (10%), and testing (10%) datasets (Table 1) in an out-of-patient manner.
Public datasets
We also used twelve public databases (1,602,960 QRS complexes from 3,050 recordings, sampling frequency from 128 to 1000 Hz) to test QRS detection performance (Table 2). Set of twelve public datasets contained both parts of public data from CinC/PhysioNet Challenge 201421, CPSC-2019 database22, CYBHi database23 using later created annotations11, European ST-T database (EDB)24, St. Petersburg INCART database (available from PhysioNet25), Lobachevski University Database (LUDB)26, MIT-BIH arrhythmia database12, QT database27, MIT-BIH ST change database (STDB)25, MIT-BIH Supraventricular Arrhythmia Database28, and T-wave alternans dataset (TWADB) from CinC/PhysioNet/Challenge 200829. These twelve databases (Table 2) were used for cross-database tests in the QRS detection task.
Databases EDB, INCART, MIT-BIH, and SVDB contained QRS classes and were used for cross-database tests of QRS classification performance (Table 2, rows highlighted with *). The proposed method is designed to classify into normal beats, PAC and PVC; therefore, if the QRS complexes were classified in more detail (e.g., a paced beat), the closest possible option was selected (e.g., a normal beat).
Training data augmentation
We randomly inverted each signal with a probability of 0.5 and cropped the signal to random 30 s, modifying the data for each batch. We used weighted oversampling to balance the counts of the QRS types we trained on.
Method
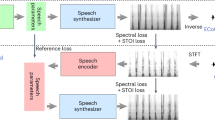
All experiments were performed in accordance with relevant guidelines and regulations. The method is designed to work as in Fig. 4. Raw ECG signal (Fig. 4A) is preprocessed (Fig. 4B), model inference (Fig. 4C) delivers probabilities of QRS classes (Fig. 4D), and post-processing (Fig. 4E) leads to resultant QRS with class information (Fig. 4F).
A workflow to detect and classify QRS with the proposed method. The ECG signal (A) is preprocessed (B) and imputted to the inference engine with the deep-learning model (C). The output of the model (D) consists of four probability vectors at the length of the input ECG signal. Output vectors are post-processed (E), and QRS annotation marks and classes are reported (F). N refers to normal sinus beat; PVC refers to premature ventricular contraction with one example of correctly detected but incorrectly classified QRS (the left-most PVC).
Preprocessing
Before feeding the training signals into the model, we resampled the signal to 100 Hz and standardised the signal independently to have zero mean and unit variance (Fig. 5A). Target data (y) for the model were prepared as follows: each QRS location was widened to 10 samples to create a four-channel segmentation mask (as in Fig. 4D).
Architecture of the presented neural network. Standardized input ECG (A) enters the first Residual Block (RB). Five of RBs are chained together (B). RB is described in further detail in (C). Then the signal enters Gated Recurrent Unit (GRU, D) layer, Fully connected layer (FC, F) and transposition (F). B × C × L refers to batch size x channel count x signal length.
Model architecture and training
The developed model consists of five residual blocks (Fig. 5B,C), a gated recurrent layer (Fig. 5D), and a fully connected layer (Fig. 5E). The model outputs a four-channel tensor (Fig. 5F). Each residual block consists of several convolutional layers. We used a batch size of 64, a cross-entropy loss function, an AdamW optimiser with a learning rate of 0.001, clipped the gradient L2 norm to 1.0, no weight decay.
Post-processing
The network outputs the likelihood (Fig. 4F) of the four different QRS classes (no QRS, normal QRS, atrial QRS, ventricular QRS) for every input sample. We take the class with the maximum likelihood for every sample and post-process the resulting segmentation mask to get a list of the QRS peaks. First, we calculate the centers of the segmentation mask and save them into a list of potential peaks. Then, we remove lower peaks that are too close (< 0.15 s) to stronger peaks, as such a low distance between beats is physiologically improbable.
Compared QRS detectors and used metrics
For comparison, we also evaluated used datasets by several publicly available QRS detection methods: by Elgendi2, Malik et al.3, XQRS detector from Python WFDB package30, and by Pan and Tompkins1. We also evaluated three-detectors from the python NeuroKit package—modified Engelse & Zeelenberg6, Hamilton detector4, and Kalidash detector7. Results for these detectors might differ from the performance reported by respective papers since we used all available data from all datasets; we implemented detectors by Elgendi2 and Malik3 using respective papers.
We used the F1 score to compare and evaluate results. A detected QRS complex was considered true positive when its annotation mark was closer than 0.1 s (inclusive) to an annotation mark prepared by an expert. The false positive or false negative cases were considered when a beat was missing in expert annotations or detected QRS complexes.
Results
The model was built using the PyTorch framework31 and trained in 70 epochs using the private MDT dataset. We separately evaluated QRS detection performance and QRS classification performance; we also evaluated computational method performance.
QRS detection performance
We received training, validation, and testing F1 scores of 0.991, 0.990, and 0.992 for the detection task using the MDT test set. We also provided a cross-database test to evaluate detection performance on twelve public datasets, showing a mean F1 score of 0.96 ± 0.06. The detection performance was compared to other methods using all twelve test datasets. We received a maximal mean F1 score of 0.961 using the proposed method, followed by the Malik method3 (0.955) and XQRS detector from the WFDB30 Python package. We also observed how the used databases were difficult for tested detectors. Overall F1 results by all detectors per database showed that the easiest database to detect was the STDB25 (0.979), the first part of PhysioNet/CinC challenge 201421 (0.972) and the MITDB (0.958). On the other hand, the most challenging database for tested detectors was the second part of PhysioNet/CinC challenge 2014 (0.759), the SVDB (0.805), and the MDT test set (0.809).
QRS classification performance
We evaluated the proposed method to classify QRS complexes into three groups—normal beat, premature ventricular contraction, and premature atrial contraction (Table 4—the last row). We reached an overall classification F1 performance of 0.96 and 0.74 for micro and macro computation in the MDT test set, respectively. Cross-database tests for QRS classification (Table 4, the first four rows) showed average micro and macro F1 scores of 0.95 ± 0.03 and 0.73 ± 0.06, respectively.
Method computational performance
We measured the processing time of all compared methods using all testing datasets (excluding the CYBHi) to evaluate computational performance. The average processing time per record is shown in Fig. 7. The comparison was obtained using a computer with Intel® Xeon® Gold 6248R CPU running at 3.00 GHz. Data were supplied to algorithms one by one, and we disabled GPU, which disadvantaged the proposed method (Fig. 7).
Discussion
The presented method showed the highest overall QRS detection F1 score in compared methods (Table 3) when using all test datasets. We were generally focused on Holter ECG data acquired during usual daily activities, and we received the best score of tested methods in the MDT dataset. The highest overall score might reflect that we used a high amount of disrupted ECG data for training. Table 3 (the row "MDT") demonstrates how different methods can detect QRS in noisy data. Figure 6 shows examples of non-trivial Holter ECG and results of presented and compared detection methods. Figure 6A demonstrates that four methods overlooked PVCs with abnormally low amplitude; Fig. 6B shows how methods react to signal disturbance and how most of them capture noise instead of QRS if they are very close (19th second). Finally, Fig. 6-C demonstrates how non-usual PVC couplets and noise may confuse detectors.
Non-trivial Holter ECG examples (parts of 30-s blocks) with QRS detection results by several methods. (A) Normal sinus rhythm with multifocal premature ventricular beats; (B) normal sinus rhythm, heavily disturbed; (C) normal sinus rhythm, disturbed, with premature ventricular beats. Bold black rectangles refer to regular QRS, filled triangles refer to premature ventricular contraction (PVC). Bold vertical at approx. second 1.3 refers to a questionable PVC combined with an artifact. Shown signals were not used for development nor testing of the presented method.
We also compared the presented method to the deep-learning method11 trained on the CYBHi dataset23 and tested on MIT-BIH12 dataset with an F1 score of 0.96. Our method slightly outperforms the compared method on MIT-BIH, but on the other hand, we used a significantly more complex network structure.
The important benefit of the presented method is that it classifies QRS complexes into three groups. Our results show that the weakest point of QRS classification is in the PAC class (Table 4). Further investigation revealed that in most cases, false PACs are generated inside blocks of atrial fibrillation where the presented method tends to report PACs. We also found incorrect classifications in long SVT runs (series of PACs running on high heart rate).
A limitation in comparison to most other methods is processing time, as shown in Fig. 7. However, this can be overpassed when the model uses a GPU during inference. In such a case, inference time can be decreased approximately 10–30 times depending on the specific hardware and batch size.
Conclusion
We presented a novel deep learning method for QRS detection and classification in one inference step. The method was evaluated on twelve public datasets not used for model development. This cross-database test showed higher overall QRS detection performance than other compared methods. Furthermore, we showed how compared QRS detectors behave in non-trivial situations common in Holter ECG. We also demonstrated that both QRS classification and detection could be combined into one deep-learning model. Therefore, the usual processing chain to analyse heart rhythm can be simplified.
Data availability
Private data supporting this study's findings are available from Medical Data Transfer, s.r.o. (Brno, Czech Republic) but they are not publicly available. However, data are available from the authors upon reasonable request and with the permission of the Medical Data Transfer company.
References
Pan, J. & Tompkins, W. J. A real-time QRS detection algorithm. IEEE Trans. Biomed. Eng. 32(3), 230–236 (1985).
Elgendi, M. Fast QRS detection with an optimised knowledge-based method: Evaluation on 11 standard ECG databases. PLoS ONE 8(9), e73557 (2013).
Malik, J., Soliman, E. Z. & Wu, H. T. An adaptive QRS detection algorithm for ultra-long-term ECG recordings. J. Electrocardiol. 60, 165–171 (2020).
Hamilton, P. & Limited, E. P. Open source ECG analysis. (2002).
Engelse, W. A. & Zeelenberg, C. A single scan algorithm for QRS detection and feature extraction. IEEE Comput. Cardiol. 2, 37–42 (1979).
A. Lourenço, H. Silva, P. Leite, R. Lourenço, and A. Fred, "REAL TIME ELECTROCARDIOGRAM SEGMENTATION FOR FINGER BASED ECG BIOMETRICS," in Proceedings of the International Conference on Bio-inspired Systems and Signal Processing, 2012, pp. 49–54.
V. Kalidas and L. Tamil, "Real-time QRS detector using stationary wavelet transform for automated ECG analysis," in Proceedings - 2017 IEEE 17th International Conference on Bioinformatics and Bioengineering, BIBE 2017, 2017, vol. 2018-January, pp. 457–461.
Mehta, S. S., Shete, D. A., Lingayat, N. S. & Chouhan, V. S. K-means algorithm for the detection and delineation of QRS-complexes in electrocardiogram. IRBM 31(1), 48–54 (2010).
Saini, I., Singh, D. & Khosla, A. QRS detection using K-Nearest Neighbor algorithm (KNN) and evaluation on standard ECG databases. J. Adv. Res. 4(4), 331–344 (2013).
Cai, W. & Hu, D. QRS complex detection using novel deep learning neural networks. IEEE Access 8, 97082–97089 (2020).
Silva, P. et al. Towards better heartbeat segmentation with deep learning classification. Sci. Rep. 10, 1–13 (2020).
Moody, G. B. & Mark, R. G. The impact of the MIT-BIH arrhythmia database. IEEE Eng. Med. Biol. Mag. 20(3), 45–50 (2001).
De Chazal, P., O’Dwyer, M. & Reilly, R. B. Automatic classification of heartbeats using ECG morphology and heartbeat interval features. IEEE Trans. Biomed. Eng. 51(7), 1196–1206 (2004).
Nascimento, N. M. M. et al. Heart arrhythmia classification based on statistical moments and structural co-occurrence. Circuits Syst. Signal Process. 39(2), 631–650 (2020).
Hammad, M., Iliyasu, A. M., Subasi, A., Ho, E. S. L. & El-Latif, A. A. A. A multitier deep learning model for arrhythmia detection. IEEE Trans. Instrum. Meas. 70, 63 (2021).
Wang, R., Fan, J. & Li, Y. Deep multi-scale fusion neural network for multi-class arrhythmia detection. IEEE J. Biomed. Heal. Informatics 24(9), 2461–2472 (2020).
Ferretti, J., Randazzo, V., Cirrincione, G. & Pasero, E. 1-D convolutional neural network for ECG arrhythmia classification. In Smart Innovation, Systems and Technologies Vol. 184 269–279 (Springer, 2021).
Oh, S. L., Ng, E. Y. K., Tan, R. S. & Acharya, U. R. Automated diagnosis of arrhythmia using combination of CNN and LSTM techniques with variable length heart beats. Comput. Biol. Med. 102, 278–287 (2018).
F. Murat, O. Yildirim, M. Talo, U. B. Baloglu, Y. Demir, and U. R. Acharya, "Application of deep learning techniques for heartbeats detection using ECG signals-analysis and review," Computers in Biology and Medicine, vol. 120. Elsevier Ltd, 01-May-2020.
Chen, A. et al. Multi-information fusion neural networks for arrhythmia automatic detection. Comput. Methods Programs Biomed. 193, 2 (2020).
Moody, G., Moody, B. & Silva, I. Robust detection of heart beats in multimodal data: The physionet/computing in cardiology challenge 2014. Comput. Cardiol. 41, 549–552 (2014).
Gao, H. et al. An open-access ECG database for algorithm evaluation of QRS detection and heart rate estimation. J. Med. Imaging Heal. Inf. 9, 9 (2019).
da Silva, H. P., Lourenço, A., Fred, A., Raposo, N. & Aires-de-Sousa, M. Check Your Biosignals Here: A new dataset for off-the-person ECG biometrics. Comput. Methods Programs Biomed. 113(2), 503–514 (2014).
Taddei, A. et al. The European ST-T database: Standard for evaluating systems for the analysis of ST-T changes in ambulatory electrocardiography. Eur. Heart J. 13, 9 (1992).
Goldberger, A. L. et al. PhysioBank, PhysioToolkit, and PhysioNet: components of a new research resource for complex physiologic signals. Circulation 101(23), E215–E220 (2000).
Kalyakulina, A.I. et al., "LU electrocardiography database: A new open-access validation tool for delineation algorithms," arXiv. 2018.
Laguna, P., Mark, R. G., Goldberg, A. & Moody, G. B. Database for evaluation of algorithms for measurement of QT and other waveform intervals in the ECG. Comput. Cardiol. 2, 5 (1997).
Greenwald, S. D., Patil, R. S. & Mark, R. G. Improved detection and classification of arrhythmias in noise-corrupted electrocardiograms using contextual information within an expert system. Biomed. Instrum. Technol. 26, 2 (1992).
Moody, G. B. The physionet/computers in cardiology challenge 2008: T-wave Alternans. Comput. Cardiol. 35, 25 (2008).
Silva, I. & Moody, G. B. An open-source toolbox for analysing and processing physionet databases in MATLAB and octave. J. Open Res. Softw. 2, 2 (2014).
Paszke, A. et al. PyTorch: An imperative style, high-performance deep learning library. Adv. Neural Inf. Process. Syst. 32, 2 (2019).
Acknowledgements
This work was supported by the Czech Technological Agency grant number FW01010305 and the CAS project RVO:68081731.
Author information
Authors and Affiliations
Contributions
A.I. – method development and evaluation, writing the manuscript I.V. – method validation P.N. – method development R.S. – annotating data Z.K. – method validation V.B. – data management J.H. – method evaluation P.J. – writing the manuscript F.P. – writing the manuscript; developing a software tool for annotating data; annotating data.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ivora, A., Viscor, I., Nejedly, P. et al. QRS detection and classification in Holter ECG data in one inference step. Sci Rep 12, 12641 (2022). https://doi.org/10.1038/s41598-022-16517-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-16517-4
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.