Abstract
Most of our knowledge on the ericoid mycorrhizal (ErM) symbiosis comes from temperate heathlands characterized by acidic peaty soils and many experiments with a few ascomycetous fungi. However, ericaceous plants thrive in many other ecosystems and in temperate coniferous forests, their seedlings often prosper on decomposing wood. While wood is typically exploited by basidiomycetous ectomycorrhizal (EcM) and saprobic fungi, the role of ErM fungi (ErMF) is much less clear. We explored the cultivable mycobiota of surface sterilized hair roots of Vaccinium spp. growing on decomposing wood in two coniferous forests in Mid-Norway (Scandinavia) and Northern Bohemia (Central Europe). Obtained isolates were identified using molecular tools and their symbiotic potential was tested in vitro. While the detected community lacked the archetypal ErMF Hyaloscypha hepaticicola and the incidence of dark septate endophytes and EcM fungi was negligible, it comprised other frequent asexual ascomycetous ErMF, namely H. variabilis and Oidiodendron maius, together with several isolates displaying affinities to sexual saprobic H. daedaleae and H. fuckelii. Ascomycete-suppressing media revealed representatives of the saprobic basidiomycetous genera Coprinellus, Gymnopilus, Mycena (Agaricales), and Hypochnicium (Polyporales). In the resyntheses, the tested basidiomycetes occasionally penetrated the rhizodermal cells of their hosts but never formed ericoid mycorrhizae and in many cases overgrew and killed the inoculated seedlings. In contrast, a representative of the H. daedaleae/H. fuckelii-related isolates repeatedly formed what morphologically appears as the ErM symbiosis and supported host's growth. In conclusion, while basidiomycetous saprobic fungi have a potential to colonize healthy-looking ericaceous hair roots, the mode(-s) of their functioning remain obscure. For the first time, a lineage in Hyaloscypha s. str. (corresponding to the former Hymenoscyphus ericae aggregate) where sexual saprobes are intermingled with root symbionts has been revealed, shedding new light on the ecology and evolution of these prominent ascomycetous ErMF.
Similar content being viewed by others
Introduction
Most of our knowledge on the ericoid mycorrhizal (ErM) symbiosis comes from temperate heathlands of the Northern Hemisphere that are characterized by acidic peaty soils high in recalcitrant phenolic compounds and low in available mineral nutrients and from many experiments with a few readily cultivable ascomycetous ErM fungi (ErMF), especially Hyaloscypha hepaticicola and Oidiodendron maius (Leake and Read 1991; Smith and Read 2008). It is widely accepted that under this scenario, these ErMF benefit their core Ericaceae hosts (= members of the early anther inversion clade of Ericaceae as defined in Kron et al. (2002), in the following text as ericaceous hosts, plants, etc.) mainly through improving nutrient uptake and alleviating substrate toxicity (Read and Kerley 1995; Read 1996; Perotto et al. 2002). However, ericaceous plants co-dominate vegetation in many other ecosystems (Kron and Luteyn 2005) and the spectrum of potential ErMF is much wider (Leopold 2016; Vohník 2020). In some ecosystems, ericaceous hair roots lack or are not dominated by the archetypal ErMF H. hepaticicola and O. maius and their place is taken by other mycobionts whose functioning is not fully understood (see Bruzone et al. (2015) and references therein). These often represent novel fungal lineages that occur in less explored locations (e.g., Midgley et al. 2016, 2018; Leopold et al. 2021; Vohník et al. 2022) and/or are difficult to grow in pure culture, e.g., members of Chaetothyriales (Allen et al. 2003; Lukešová et al. 2015; Toju et al. 2016; Baba and Hirose 2021), Sebacinales (Allen et al. 2003; Selosse et al. 2007; Vohník et al. 2016; Griffin and Kernaghan 2022), and Kurtia argillacea in Hymenochaetales (Kolařík and Vohník 2018). It has been suggested that some of them may confer adaptations distinct from those provisioned by the so far investigated ascomycetous ErMF, e.g., the ability to degrade recalcitrant aromatic substrates like lignin (Vohník et al. 2012a), the second most abundant biopolymer on Earth (Baucher et al. 1998).
Except confirmed and probable ErMF, ericaceous plants associate with a plethora of root mycobionts with unknown symbiotic status, including typical ectomycorrhizal (EcM) and saprobic basidiomycetes (e.g., Allen et al. 2003; Bougoure et al. 2007; Walker et al. 2011; Grelet et al. 2017). Under artificial conditions, these may form intracellular hyphal loops or pegs in the rhizodermal cells of ericaceous plants (e.g., Walker et al. 2011; Villarreal-Ruiz et al. 2012; Vohník et al. 2012a; Grelet et al. 2017) and even support the growth of the inoculated plants (Vohník et al. 2012b; Grelet et al. 2017), but the eco-physiological significance of such observations remains unclear. The mechanisms behind these positive effects are unknown and may include a release of nutrients to the host´s rhizosphere through autolysis of their mycelium (Duclos et al. 1983) and mineralization of organically bound nutrients (Vohník et al. 2012b; Grelet et al. 2017). In addition, ericaceous plants often associate with the so-called dark septate endophytes (DSE), a miscellaneous group of ascomycetous mycobionts with melanized hyphae that are ubiquitous in the roots of boreal and temperate plants and whose effects range from positive to negative (Newsham 2011; Mayerhofer et al. 2013). DSE may form intracellular hyphal coils in the rhizodermis of ericaceous roots (Massicotte et al. 2005; Vohník and Albrechtová 2011; Lukešová et al. 2015) and similarly to non-ericaceous hosts, their effects range from slightly positive to neutral to negative (Vohník et al. 2003, 2005; Lukešová et al. 2015).
Ericaceous plants are not limited to acidic peaty substrates and in many European boreal and temperate forests, they often grow in soil mixtures comprising various volumes of decomposing wood (Fig. 1). Decaying tree stumps and thick branches laying on the forest floor seem to be especially suitable for the otherwise rare European blueberry (Vaccinium myrtillus) seedlings (Welch et al. 2000) as they retain moisture and provide elevated surfaces that assure an escape from competition with other forest floor plants (M. Vohník, personal observations). Dead wood forms a large part of the total biomass not only in boreal and temperate forests and represents an important pool of organically bound mineral nutrients (especially nitrogen and phosphorus) that are, however, directly non-accessible to primary producers and have to be released by heterotrophs, primarily basidiomycetous and to a lesser extent ascomycetous fungi (Boddy and Watkinson 1995). The major components of wood are cellulose, lignin and hemicelluloses and while many fungi are cellulolytic, they often cannot access lignin (Janusz et al. 2017; Goodell et al. 2020). Despite that the enzymatic repertoire of the so far investigated ascomycetous ErMF comprises enzymes involved in plant cell wall degradation (Cairney and Burke 1998; Perotto et al. 2018; Martino et al. 2018), their ability to degrade cellulose and especially lignin seems to be lower relative to the wood decomposing basidiomycetous saprobic fungi (Pearson and Read 1975; Bending and Read 1997). In contrast, K. argillacea, the basidiomycetous mycobiont forming sheathed ericoid mycorrhiza (a morphotype of ericoid mycorrhiza where intracellular hyphal coils in the host rhizodermis are accompanied by often multiple layers of thick hyphae with clamp connections on the root surface), has both cellulolytic and ligninolytic abilities (Vohník et al. 2012a). In addition, some Australian ErMF may outperform H. hepaticicola in utilization of phenolic compounds (Midgley et al. 2006).
European blueberry (Vaccinium myrtillus) growing on decomposing wood, an overlooked substrate regularly colonized by many temperate ericaceous species. Next to nothing is known about the diversity of the fungi colonizing ericaceous hair roots growing in this substrate and the role these fungi play in wood decomposition. The picture was taken at the North Bohemian site investigated in this study (see Materials and methods), the peak in the background is Luž (Lausche in German, Łysa in Sorbian), the highest peak (793 m a. s. l.) of the Lusatian Mountains located at the border between Czechia and Germany
To our knowledge, the root mycobiota of ericaceous plants growing on decomposing wood has not been investigated. Therefore, here we present the results of two surveys from Mid-Norway (Scandinavia) and Northern Bohemia (Central Europe) focused on fungi associated with Vaccinium spp. hair roots in this overlooked substrate. Both surveys were mainly observational, i.e., we did not rigorously test any specific hypotheses. On the other hand, thanks to the specific nature of decomposing wood, especially when compared to peat, we expected that the screened hair roots would harbor a spectrum of mycobionts differing from those commonly encountered in peatland ecosystems, possibly enriched in typical basidiomycetous EcM and saprobic fungi. The surveys were accompanied by microscopic observations of the root colonization and two in vitro resynthesis experiments and their results are presented together with the diversity and phylogenetic data.
Materials and methods
Sampling
A first set of samples was collected in May 2011 in a naturally regenerating Norway spruce (Picea abies) forest at the foothill of Forbordfjellet (N63.51877, E10.88811; ca. 400 m above sea level) close to Stjørdal in Mid-Norway. Mixed roots of European blueberry (Vaccinium myrtillus) and cowberry (V. vitis-idaea) overgrowing a partially decomposed spruce stump were collected and processed as described in Vohník et al. (2012a) with a special focus on evaluating the presence of sheathed ericoid mycorrhiza and isolating basidiomycetous root-associated fungi. A second set was collected in June 2011 in a secondary Norway spruce forest in Northern Bohemia, Czechia (N50.85593, E14.61735; ca. 664 m a. s. l.; in 2022, all adult spruce trees were cut down). Roots of 12 ca. 3-year-old European blueberry seedlings growing on decomposing thick branches in four slash piles left after a thinning were collected and processed in the same manner as above with a focus on describing their fungal colonization and isolating their ascomycetous and basidiomycetous mycobionts.
Herbarium specimens (as dried cultures) were deposited in the Herbarium of the Institute of Botany Institute, Czech Academy of Sciences in Průhonice (PRA), cultures were accessioned into Westerdijk Fungal Biodiversity Institute in Utrecht, the Netherlands (CBS).
Microscopic observations of natural colonization
The root samples were washed under running tap water, ca. one half of both sets was cleared with 10% KOH for 15 min at 121 °C, briefly acidified with 3% HCl and washed under running tap water. The roots from Northern Bohemia were further stained with 0.05% trypan blue in lactoglycerol (glycerol + lactic acid + water, volume ratio 2:1:2) overnight and de-stained in lactoglycerol. Microscopic observations of the roots were performed with a compound Olympus BX60 microscope equipped with differential interference contrast at 400 × and 1000 × magnification. Photos were taken with an Olympus DP70 camera using QuickPHOTO MICRO v. 3.2 (Promicra) and the embedded Deep Focus Mode was employed when needed. The obtained photos were modified for clarity and contrast as needed and assembled into figures using Paint.net v. 4.0.13 (dotPDN LLC, Rick Brewster and contributors).
Isolation of mycobionts
The remaining washed roots were surface sterilized for 30 s in 10% SAVO (common household bleach; Unilever; 100% SAVO contains 47 g/kg, i.e., 4.7%, sodium hypochlorite, NaClO), 3 × washed with sterile deionized water and ca. 2.5 mm segments of hair roots were plated on the surface of a growth medium [modified Melin Norkrans medium (MMN); Molina and Palmer (1982)] amended with Novobiocin sodium salt (50 mg/l; Sigma-Aldrich) to prevent growth of bacteria and incubated in the dark at room temperature for ca. two months. The segments from Northern Bohemia were cultivated on MMN with and without benomyl (4 mg/l, reduces growth of most ascomycetes; Sigma-Aldrich) whereas the segments from Mid-Norway were incubated only on MMN with benomyl. Sporulating mycelia were discarded and those non-sporulating were transferred to new dishes with MMN and used for molecular identification.
DNA extraction, PCR amplification and sequencing
DNA was extracted from all isolates using an Extract-N-Amp Plant Kit (Sigma-Aldrich) following the manufacturer’s instructions. The ITS1-5.8S-ITS2 nuclear ribosomal DNA (rDNA) region was amplified using the ITS1F + ITS4 primer pair, with PCR parameters and gel electrophoresis as described in Vohník et al. (2012a). The PCR products were purified and sequenced in Macrogen Europe Laboratory (Macrogen) using the ITS1, ITS1F and ITS4 primers.
Identification of mycobionts
The obtained sequences were screened in Finch TV v. 1.4.0 (Geospiza) and manually edited. Subsequently, the Czech and the Norwegian ones were separately aligned in Bioedit v. 7.1.8 (Hall 1999) and clustered at 99% similarity in TOPALi (Biomathematiscs & Statistics Scotland) (Tables 1 and 2). Representative sequences of each cluster were subjected to BLASTn searches (Zhang et al. 2000) in GenBank at NCBI (Sayers et al. 2019) as detailed in (Vohník 2020). Sequences of several Czech isolates displayed affinities to Hyaloscypha s. str. but did not seem to belong to any of the so far described and sequenced species. The amplified ITS rDNA of their representatives was sequenced with the ITS4 primer and LSU rDNA amplified with the LR0R + LR5 primer pair and sequenced using the same primers in Macrogen Europe Laboratory. Raw sequence data of these new hyaloscyphoid isolates were assembled and edited using Sequencher v. 5.4.6 (Gene Codes).
Phylogenetic analyses
Since the new hyaloscyphoid isolates were sterile in culture and no diagnostic features were detected, their closest relatives were selected from the top ranked hits using BLASTn searches of the ITS and LSU sequences generated in this study. The homologous ITS and LSU sequences of representatives of Hyaloscyphaceae were retrieved from GenBank at NCBI. The GenBank accession numbers of all strains analyzed are listed in Table 3.
The ITS and LSU sequences were aligned in Mafft v. 7.487 (Katoh and Standley 2013) implemented in the CIPRES Science Gateway v. 3.3 (Miller et al. 2010) and manually corrected in Bioedit v. 7.1.8 (Hall 1999) when necessary. Single-locus ITS and LSU data sets for representatives of Hyaloscyphaceae (ITS: 52 sequences/536 characters including gaps, LSU: 41/1302), for which we assumed rate heterogeneity, were evaluated using MrModeltest v. 2.4 (Nylander 2004) to find the best partitioning scheme and to select best-fit models under the corrected Akaike information criteria. The SYM + G best-fit model was selected for both partitions. The concatenated ITS-LSU dataset (deposited in TreeBase 29,685) was subjected to phylogenetic analyses. The first 89 nucleotides of LSU at the 5′-end were excluded from the alignment because of the incompleteness in most sequences. The dataset consisted of 1838 characters including gaps and 343 unique character sites. Three members of the genus Hyphodiscus (Hy.) (Hyphodiscaceae, Helotiales), namely Hy. brachyconius, Hy. brevicollaris and Hy. luxurians, were used to root the tree.
Phylogenetic relationships were evaluated using maximum likelihood (ML) and Bayesian Inference (BI) analyses and were performed through the CIPRES Science Gateway v. 3.3. ML analysis was performed with RAxML-HPC v. 8.2.12 (Stamatakis 2014) with a GTRCAT approximation. Nodal support was determined by non-parametric bootstrapping (BS) with 1000 replicates. BI analysis was performed in a likelihood framework as implemented in MrBayes v. 3.2.6 (Huelsenbeck and Ronquist 2001) using default parameters. The B-MCMCMC analysis lasted until the average standard deviation of split frequencies was below 0.01 with trees saved every 1000 generations. The first 25% of saved trees, representing the burn-in phase of the analysis, were discarded. The remaining trees were used for calculating posterior probabilities (PP) of recovered branches. Obtained trees were viewed in SeaView v. 4 (Gouy et al. 2010) and edited in MS PowerPoint (Microsoft).
First resynthesis
Mycelial cultures of five fungal isolates from Northern Bohemia (three ascomycetes and two basidiomycetes, Table 1) were pre-cultivated in Petri dishes on potato carrot agar (PCA) in the dark at room temperature for six weeks. Seeds of European blueberry of local origin were extracted from fresh fruits, surface sterilized 1 min in 10% SAVO, 3 × washed with sterile deionized water and left to germinate and develop on MMN with no maltose, 1 g/L glucose and 50 μg/L Novobiocin added to suppress possible bacterial growth in a growth chamber under a 21 °C – 16 h light/15 °C – 8 h dark cycle and irradiation of 200 μmol/m2 s1 for 3 months. The cultivation substrate consisted of peat, perlite, and a mixture of dead wood of local origin (extracted from an old Norway spruce stump in the Průhonice Park) and wood shavings (volume ratio 4:2:1). The substrate was added to Magenta GA-7 cultivation vessels (Sigma-Aldrich), 25 g per each vessel, and watered with 20 mL of sterile deionized water. The vessels with the substrate were double autoclaved (60 min at 121 °C repeated after 36 h) and when cooled down, plugs (diam. ca. 5 mm) excised from the mycelial cultures were added on the surface of the substrate (four plugs per each vessel). A control treatment was established with PCA plugs without fungal mycelium. Each vessel was watered with 10 mL of sterile deionized water, the plugs were mixed into the substrate and one blueberry seedling was inserted into the substrate per each vessel. There were five vessels per treatment, incl. the fungus-free control. The vessels were incubated in the same growth chamber under the same regime as above and periodically checked. The experiment was harvested after 137 days, the seedlings were gently removed from the substrate and their shoots were separated from the roots. The shoots were dried at 65 °C overnight and weighed. The roots were gently washed under running tap water to remove residues of the cultivation substrate and treated as described above for the naturally colonized roots, including microscopy. Three small pieces of the cultivation substrate were extracted from each vessel and placed on the surface of solidified malt extract agar (MEA, HiMedia) in plastic Petri dishes (9 cm in diam.) and incubated in the dark for 1 month. DNA was extracted from representative cultures and ITS rDNA was amplified and sequenced as described above. The obtained sequences were identified as described above and compared with the sequences of the inoculated fungi.
Second resynthesis
Mycelial cultures of seven fungal isolates, three from Northern Bohemia and four from Mid-Norway (Tables 1 and 2), one ascomycete and six basidiomycetes, were pre-cultivated as above. The cultivation substrate consisted of peat and perlite (volume ratio 1:1) and 14 g of the substrate were added to each Magenta GA-7 vessel. The vessels were double autoclaved as above except that before the second autoclaving, 50 mL of molten 0.8% water agar amended with 0.1% active charcoal were pipetted over the substrate in each vessel. After cooling down, a small piece of substrate was extracted from 10 random vessels and the pieces were aseptically transferred to plastic Petri dishes (5 cm in diam.) with MEA. The vessels and the dishes were incubated for 2 weeks at room temperature in the dark to doublecheck their sterility and no fungal mycelium was detected upon final inspection. Subsequently, plugs (diam. ca. 3 mm) excised from the mycelial cultures were added on the surface of the substrate (nine plugs per each vessel), a control treatment obtaining PCA plugs without fungal mycelium. The vessels with the plugs were incubated as above and after two months, all plugs extracted from fungal cultures were covered with mycelium spreading over the surface of the cultivation substrate (control plugs did not produce any mycelium). The plugs were mixed into the substrate and one 3-months-old European blueberry seedling obtained as above was inserted into the substrate per each vessel. There were five vessels per treatment, incl. the fungus-free control. The vessels were incubated in the same growth chamber under the same regime as above and periodically checked. The experiment was harvested after 158 days and the seedlings were treated as described above.
Statistical analyses
Because the datasets from the two resynthesis experiments violated the assumptions of ANOVA, the effects of inoculation on dry shoot weight (both resyntheses) and fresh root weight (second resynthesis) were evaluated using the non-parametric Kruskal–Wallis test followed by multiple comparisons of mean ranks (Dunn´s test) in STATISTICA v. 12 (Statsoft).
Results
Microscopic observations of natural colonization
Sheathed ericoid mycorrhizae were found only in the samples from Mid-Norway. In both sets of samples, the hair roots displayed typical ericoid mycorrhizal colonization, i.e., dense intracellular hyphal coils in the rhizodermis. The colonization often started already at the tips of the hair roots (Fig. 2A) and in the samples from Northern Bohemia, it was often accompanied by extensive extraradical mycelium (Fig. 2B). In addition, many hair roots from Northern Bohemia were covered by sparse mantles formed by thin, often undulated hyphae that were connected with the dense intracellular hyphal coils in the rhizodermis (Fig. 2C and D). Both sets of samples contained thick, often interwoven hyphae with clamp connections, but these were never seen penetrating the screened roots (now shown). Unexpectedly, in both sets of samples there were no melanized microsclerotia typical of the dark septate endophytes. While the Czech samples also lacked thick melanized surface hyphae, these were regularly present in the Norwegian samples and in many cases, they seemed to be directly connected with much thinner intracellular hyphae occurring in the host´s rhizodermis and morphologically resembling ericoid mycorrhizae (Fig. 3). Sometimes, such connections seemed to be realized through haustoria-like intracellular structures (Fig. 3C‒G) that are, to our best knowledge, not known from resynthesis experiments with typical ErMF.
Natural fungal colonization in hair roots of Vaccinium myrtillus from Northern Bohemia. A: Typical ericoid mycorrhizal pattern, i.e., intracellular hyphal coils, present in most of the hair root´s rhizodermal cells (arrows point at some examples). Only a few rhizodermal cells remain without fungal colonization (examples marked with asterisks). B: Extraradical mycelium (arrow) accompanying ericoid mycorrhizal colonization (examples marked with asterisks). C, D: Sparse mantles formed by thin, often undulated hyphae on the surface of the hair root (arrows). Ericoid mycorrhizal colonization occurs just below the nets, in the host´s rhizodermis (examples marked with asterisks). All roots were cleared with 10% KOH, stained with trypan blue and observed using a compound microscope equipped with differential interference contrast as described in Materials and Methods. Scale bars = 20 μm
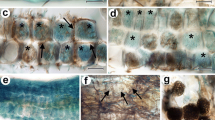
Natural fungal colonization in hair roots of Vaccinium spp. from Mid-Norway. Eight consecutive views of a hair root with dark septate hyphae on the surface (open arrows point at some examples in A) and intracellular hyphal colonization morphologically corresponding to ericoid mycorrhiza (examples in A and B marked with asterisks). C–F: An intracellular haustorium-like structure (closed arrows) is connected with a dark septate hypha on the surface through a thin penetration hypha (closed arrowheads in C–H). Note that the same rhizodermal cell is also occupied with dense hyphal coils (A, B). A–H: Another dark septate hypha on the surface is connected through a penetration hypha (open arrowheads) with intracellular hyphal colonization corresponding to ericoid mycorrhiza. The root was cleared with 10% KOH and observed using a compound microscope equipped with differential interference contrast as described in Materials and Methods. Scale bars = 20 μm
Isolation and identification of mycobionts
The Czech samples yielded 65 isolates with high-quality sequences that were grouped into 16 OTU at 99% sequence similarity. The most abundant was Hyaloscypha variabilis (19 isolates) followed by Hyaloscypha sp. (10), Pezicula sp. (10), Oidiodendron maius (6), etc. Nine OTU were represented only by one isolate (Table 1). The Norwegian samples yielded 27 isolates grouped into 8 OTU and the most abundant was Mycena sp. (17). The three following OTU belonged to the same genus and in total comprised 6 isolates. Two OTU with a single isolate belonged to the genus Gymnopilus, one to Hypochnicium and one to Serendipita (Table 2).
Phylogenetic analyses
Relationships of the new hyaloscyphoid isolates B3, B23, B58, B62, B67, B80, and CBS 149189 (see Hyaloscypha sp. above) obtained from hair root of Vaccinium myrtillus from Northern Bohemia were assessed in the phylogenetic analysis based on ITS-LSU sequences. Sequences of 43 strains representing 21 species of the genus Hyaloscypha s. str. available in GenBank were included in the analysis, as well as two unnamed fungal isolates that showed the highest conformity in the BLASTn searches. The ML and BI trees were highly concordant; the ML tree is shown in Fig. 4. The new hyaloscyphoid isolates clustered into a strongly supported clade consisting of H. daedaleae, H. leuconica, H. fuckelii and the two unnamed fungal isolates. The isolates B3 and B23 formed a monophyletic subclade while the rest of the isolates formed a separate lineage with little differentiation among them. These two groups differ slightly in their ITS sequences. The isolates B3 and B23 have identical ITS sequences and the isolates that clustered around the isolate B10 (CBS 149189) show 99.79–100% sequence identity, while ITS sequence identity between the two groups ranges from 98.75 to 98.95%. Based on this comparison and position in the phylogenetic tree, it is likely that they represent two different species. The closest named relatives to the two groups of the new hyaloscyphoid isolates are two strains of H. fuckelii (AMFB1780 and TK7053) and three strains of H. daedaleae (CBS 120.91, CBS 120.92, ZW-Geo138-Clark).
Maximum likelihood tree based on combined ITS and LSU rDNA sequences of members of Hyaloscypha s. str., showing the phylogenetic position of the new hyaloscyphoid isolates. Abbreviations T and IT after the name indicate ex-type and ex-isotype strains, respectively. Species names placed in grey boxes have been linked with mycorrhizal or root-endophytic lifestyle. Species names placed in the light blue box indicate closest relatives to our hyaloscyphoid isolates, which are placed in the blue box. The arrow points out the position of Hyaloscypha fuckelii CBS 126292, which is distant from H. fuckelii AMFB11780 and H. fuckelii TK7053 in the light blue box. Thickened branches indicate branch support with ML BS = 100% and PP values = 1.0. Branch support of nodes ≥ 75% ML and ≥ 0.95 PP is indicated above or below branches
First resynthesis
Compared to the non-inoculated control seedlings, those inoculated with H.yaloscypha variabilis B78, Hyaloscypha sp. CBS 149189 (B10) and O. maius B56 grew well, had green to green–brown leaves and did not show any symptoms of stress. In contrast, the control seedlings’ leaves were smaller and typically yellowish to reddish. Similar was true for the seedlings inoculated with Mycena sp. B14. The seedlings inoculated with Gymnopilus sp. B83 were overgrown by whitish mycelium and three of them died, hence they were excluded from the statistical analysis (see below). All pieces of the substrate collected for a verification of the inoculation success produced mycelium with ITS rDNA sequences matching those of the inoculated fungi, except the substrate inoculated with Mycena sp. B14 that produced no mycelium. The control substrate produced no mycelium.
Hyaloscypha variabilis B78 and O. maius B56 colonized the roots of all inoculated seedlings but their colonization patterns did not resemble ericoid mycorrhiza (Fig. 5A). Similarly, the roots of the two surviving seedlings inoculated with Gymnopilus sp. B83 displayed locally abundant fungal colonization, including intracellular hyphae in the rhizodermis, but distinct from the typical ErM colonization pattern (Fig. 5B). Only Hyaloscypha sp. B10 CBS 149189 formed what morphologically corresponds to the ErM symbiosis (see below). The roots of the seedlings inoculated with Mycena sp. B14 were free of any fungal colonization, and therefore, this treatment was excluded from the statistical analysis. Roots of the non-inoculated control seedlings did not display any signs of fungal colonization.
Results of the two resyntheses with Vaccinium myrtillus seedlings. A: Thin intracellular fungal hyphae resembling endophytic colonization rather than ericoid mycorrhiza (arrows point at some examples). Many rhizodermal cells are free of fungal colonization (asterisks mark some examples), the presence of intact nuclei (arrowheads point at some examples) suggests that the cells were metabolically active. First resynthesis, Hyaloscypha variabilis B78. B: Heavy fungal colonization at the tip of a hair root. Some cells are filled with fungal hyphae (arrows point at some examples) and this pattern resembles endophytic colonization rather than ericoid mycorrhiza. First resynthesis, Gymnopilus sp. B83. C: Typical ericoid mycorrhizal colonization by Hyaloscypha sp. B10 in contrast with non-colonized rhizodermal cells (asterisks mark some examples). Second resynthesis. D: Rhizodermal cells packed with dense hyphal coils typical for ericoid mycorrhiza (asterisks mark some examples). Note that while there are some signs of fungal colonization in the outer cortex (arrows), fungal hyphae never penetrate the stele/vascular tissues. Second resynthesis, Hyaloscypha sp. B10. E: Typical ericoid mycorrhizal colonization by Hyaloscypha sp. B10. Note that while nearly all rhizodermal cells are colonized (asterisks mark some examples), the cells below the rhizodermis do not show signs of fungal colonization. F: Another example of ericoid mycorrhizal colonization by Hyaloscypha sp. B10. Note that while all rhizodermal cells are colonized (asterisks), the fungal colonization extends also to the cells below the rhizodermis (arrows). All roots were cleared with 10% KOH, stained with trypan blue and observed using a compound microscope equipped with differential interference contrast as described in Materials and Methods. Scale bars = 20 μm
Second resynthesis
Only the seedlings inoculated with Hyaloscypha sp. B10 prospered better than the non-inoculated control and produced green to green–brown leaves. The substrate in all vessels inoculated with the basidiomycetes was colonized by whitish mycelia and in the vessels inoculated with Gymnopilus sp. B83, Mycena sp. JPK-117 and Mycena sp. JPK-152, they overgrew and subsequently killed 2–3 seedlings per treatment; consequently, these treatments were excluded from the statistical analysis. Dense intracellular hyphal coils typical for the ErM symbiosis were found only in the seedlings inoculated with Hyaloscypha sp. B10 (Fig. 5C–F). There were some extraradical hyphae visible on the surface of the roots of the seedlings inoculated with Gymnopilus JPK-155, Mycena sp. B14 and Mycena sp. JPK-151, but no intracellular hyphal colonization could be seen. In the non-inoculated control treatment, no mycelium was visible in the substrate and in and around the roots.
Statistical analyses
In the first resynthesis, the inoculation significantly affected the seedlings' dry shoot weight (H = 9.789, p = 0.021) and the inoculation with Hyaloscypha sp. B10 significantly increased the seedlings' dry shoot weight in comparison with the non-inoculated control (16.62 ± 2.99 mg vs. 1.44 ± 2.99 mg, mean ± SE, p = 0.014). In the second resynthesis, the inoculation significantly affected the seedlings’ dry shoot weight (H = 12.730, p = 0.013) and fresh root weight (N = 16.634, p = 0.002). The inoculation with Hyaloscypha sp. B10 significantly increased the seedlings' dry shoot weight in comparison with the inoculation with Gymnopilus sp. JPK-155 (16.49 ± 1.30 mg vs. 1.30 ± 1.30 mg, p = 0.012) while its effect in comparison with the non-inoculated control (1.67 ± 1.30 mg) was marginally significant (p = 0.060). In comparison with the non-inoculated control, the inoculation with Hyaloscypha sp. B10 significantly increased the seedlings' fresh root weight (32.00 ± 3.22 mg vs. 1.2 ± 3.22 mg, p = 0.016).
Discussion
Some basidiomycetous ErMF and saprobes like white rot fungi have a higher lignocellulolytic potential than the prominent ascomycetous ErMF H. hepaticicola (Bending and Read 1997; Vohník et al. 2012a), so the leading idea behind this study was that these would be more abundant in substrates containing higher amounts of lignin and cellulose, namely in decomposing wood. However, this was true only in the case of the samples from Mid-Norway where we had used an ascomycete-suppressing isolation medium. In contrast, the samples from Northern Bohemia subjected to a common medium without benomyl yielded a mycobiont spectrum dominated by helotialean ascomycetes, as is common for ericaceous plants dominating the archetypal peatland ecosystems of the Northern Hemisphere. While the resyntheses with saprobic basidiomycetes from Mid-Norway and Northern Bohemia brought at best inconclusive results, the Northern Bohemian samples yielded several hyaloscyphoid mycobionts with affinities to H. fuckelii sensu Kosonen et al. (2021) and H. daedaleae and their representative repeatedly formed what morphologically corresponds to the ErM symbiosis, and supporting the growth of the inoculated plants.
Microscopic observations of natural colonization
Our investigations of root mycobionts of cultured and natural populations of Vaccinium spp. in Mid-Norway had started with microscopic observations of root colonization and resulted in the discovery of a new ErM morphotype that was named sheathed ericoid mycorrhiza. Due to the difficulties met during early attempts to isolate the respective mycobiont and later to amplify its rDNA using universal primers (Vohník et al. 2012a), microscopic observations remained the easiest way how to detect sheathed ericoid mycorrhiza and its hymenochaetoid mycobiont Kurtia argillacea. Hence, this study started with microscopic observations but contrary to our expectations, we did not find the characteristic basidiomycetous morphotype in V. myrtillus seedlings from Northern Bohemia, despite that their roots only occurred in decomposing wood. On the one hand, the surfaces of these hair roots were often covered by loose hyphal mantles. On the other hand, in comparison with K. argillacea, the hyphae forming these mantles were thinner and lacked clamp connections typical for many basidiomycetous mycobionts. Such a colonization pattern was not observed in the two resyntheses and the identity of the respective mycobiont(-s) thus remains unknown. The fact that these surface hyphae lacked clamp connections does not necessarily mean that they were of an ascomycetous origin. For example, some sebacinoid mycobionts can form ericoid mycorrhizae and cavendishioid ectendomycorrhizae, both comprising hyphae occurring on the root surface, and despite being basidiomycetes, they lack clamp connections (Setaro et al. 2006; Vohník et al. 2016). This question could be at least partially answered by transmission electron microscopy focused on the anatomy of the hyphal septa, which differ between asco- and basidiomycetes (e.g., Bonfante-Fasolo 1980; Selosse et al. 2007).
DSE are very common inhabitants of ericaceous roots (e.g., Massicotte et al. 2005; Vohník and Albrechtová 2011; Gorzelak et al. 2012) and while dark septate hyphae were common on the surface of the hair roots collected in Mid-Norway, they were absent in the hair roots from Northern Bohemia. Since only one isolate belonging to the Phialocephala fortinii s. l. – Acephala applanata species complex (Grünig et al. 2008) was recovered from these roots, it appears that moist decaying wood is not a suitable substrate for these mycobionts. A similar situation has been reported for ericaceous plants growing at an acidic wetland site in SW Canada (Hambleton and Currah 1997), suggesting that water content may be a primary edaphic factor influencing the distribution of DSE associating with core Ericaceae.
Ericoid mycorrhiza and DSE association are two morphologically distinct root-fungus symbioses characterized by fine intracellular hyphal coils in the rhizodermis of the hair roots of ericaceous plants and intracellular melanized microsclerotia, respectively. However, in some cases there seems to be a morphological continuum between these two colonization patterns (Vohník and Albrechtová 2011) and the samples from Mid-Norway provided one such example, namely thick melanized surface hyphae becoming thin, hyaline and coiled upon entering the host cell. This was sometimes connected with a formation of haustoria-like intracellular structures that seemed to be connected with intracellular hyphal coils occupying the same cells. Although these structures are occasionally observed in healthy-looking naturally colonized ericaceous hair roots (M. Vohník, personal observations), to our knowledge they have never been recorded in resynthesis studies and the identity of the respective mycobiont(-s) involved thus remains unknown.
Detected fungal spectra
There was an apparent lack of H. hepaticicola in both sets of samples. While in the case of the samples from Mid-Norway this can be explained by the ascomycete-suppressing medium, the archetypal ErMF was probably absent and substituted by other mycobionts, including the ErMF H. variabilis and O.idiodendron maius and the new hyaloscyphoid fungi, in the case of the samples from Northern Bohemia. The new hyaloscyphoid fungi clustered with H. fuckelii sensu Kosonen et al. (2021) but not with H. fuckelii CBS 126292, suggesting that this species is in need of a taxonomic revision. In addition, they clustered with H. daedaleae and since these two taxa are known only as sexual saprobic morphs, it would be interesting to include them in a resynthesis experiment with an ericaceous host for comparison. In any case, previous studies have indicated a clear separation between asexual root-symbiotic and sexual saprobic Hyaloscypha s. str. spp. (Fehrer et al. 2019; Vohník et al. 2022) and our study seems to be the first case where representatives of the different reproductive and trophic morphs intermingle in one statistically supported phylogenetic clade. Additional members of the clade are two isolates from root tips of Pinus sylvestris (Fungal spp. 2.20.4G/GenBank KM068412 and 3.44.4 J/KJ649999) and except decaying wood, the clade is thus known from both conifer (Picea abies and P. sylvestris) and deciduous (Quercus robur and V. myrtillus) hosts from Belgium, Czechia, and Finland. Such a relatively broad host and distribution range is reminiscent of other asexual root symbionts in the genus Hyaloscypha s. str., namely H. variabilis (Hambleton and Sigler 2005; Vohník et al. 2013) and H. gryndleri (Vohník et al. 2022; Daghino et al. 2022).
The genus Pezicula (asexual morph Cryptosporiopsis) belongs to Dermateaceae (Helotiales) and contains more than 130 species (MycoBank, mycobank.org, accessed 11/8/2022) that often produce apothecial ascomata on the bark of temperate woody plants (Verkley 1999). Pezicula comprises both saprobes and symbionts (pathogens and endophytes, possibly also mutualists) of a wide range of hosts (Sieber 2007; Chen et al. 2016) and many species are potent producers of biologically active secondary metabolites (e.g., Stillwell et al. 1969; Fisher et al. 1984; Noble et al. 1991; Schulz et al. 1995). Isolates belonging to this genus are from time to time obtained from ericaceous roots (e.g., Verkley et al. 2003; Sigler et al. 2005; Zijlstra et al. 2005; Walker et al. 2011) and the most common species include P. brunnea, P. ericae, P. radicicola, and P. rhizophila, the first morphologically described Pezicula (Cryptosporiopsis) species from ericaceous roots (Verkley et al. 2003). However, their symbiotic status is not clear and may range from (weak) pathogenicity to mutualism. For example, due to its repeated isolations from surface-sterilized healthy roots of several ericaceous hosts, Verkley et al. (2003) regarded P. rhizophila as an endophytic fungus. Zijlstra et al. (2005) reported that Calluna vulgaris seedlings inoculated in vitro with P. rhizophila CBS 109839 showed increased nitrogen content compared to the non-inoculated control seedlings, but no information about root colonization was provided. Finally, Walker et al. (2011) reported that in their sterile resynthesis system, two P. ericae isolates formed “hyphal complexes typical for ericoid mycorrhiza” in the roots of Vaccinium uliginosum, thus demonstrating a “potential to establish ErM associations”, despite that the colonization of the roots “tended to be low” and no photo-documentation was provided. In this study, the OTU NB-03 from Northern Bohemia most likely represented P. neosporulosa and the number of its isolates equaled that of the new hyaloscyphoid fungi. Pezicula neosporulosa was described as an endophyte/parasite of Abies spp. from China and the Netherlands (Yuan and Verkley 2015) and to our knowledge, nothing is known about its functioning in ericaceous roots.
The mycobiota of the ericaceous hair roots regularly comprises a basidiomycetous component that is mainly formed by the difficult-to-cultivate sebacinoid ErMF, but also includes non-sebacinoid ErMF, various endophytes and pathogens and some typical EcM and saprobic fungi (see Vohník (2020) and references therein). While the Mid-Norwegian samples yielded one serendipitoid isolate, shown in another study to form what morphologically corresponds to the ErM symbiosis (Vohník et al. 2016), no sebacinoid fungi were isolated from the Northern Bohemian samples. Contrary to the results of, e.g., Bougoure et al. (2007) and Lorberau et al. (2017), and despite the fact that the sampling sites were in the middle of two coniferous forests, we did not obtain any EcM fungi. However, both sets of samples yielded several basidiomycetous OTU that could be linked to genera traditionally reserved for saprobes, namely Coprinellus, Gymnopilus, Hypochnicium and Mycena, the last being the most prevalent. Mycena (Mycenaceae, Agaricales) contains more than 1900 species (mycobank.org, accessed 23/8/2022) and while these are typically saprobic, they also associate with plant roots (see Harder et al. (2021) and references therein) and engage in the orchid mycorrhizal symbiosis (e.g., Ogura-Tsujita et al. 2009; Zhang et al. 2012; Lee et al. 2015). Mycena are not uncommon in ericaceous roots (see Grelet et al. (2017) and references therein) but the mode of their interactions needs to be clarified (also see below). It is well known that many fungal pathogens may have latent endophytic stages that become harmful when the host is weakened (Schulz and Boyle 2006) and that saprobes may colonize dying or already dead cells of a hair root that otherwise looks healthy (Grunewaldt-Stöcker and von Alten 2016), possibly explaining our observations of saprobic basidiomycetes in Vaccinium spp. hair roots.
Resyntheses
Our at best inconclusive results in the resyntheses with saprobic basidiomycetes do not support previous observations of their beneficial effects on ericaceous hosts (Grelet et al. 2017). However, under certain scenarios, they can be beneficial for the growth of ericaceous plants even without forming a root-fungus symbiosis (Vohník 2020). For example, under natural conditions they never interact with fungus-free roots and there is an indication that they might benefit ericaceous plants through interactions with ErMF (Vohník et al. 2012b). More experimental work is apparently needed to resolve this issue, ideally employing a combined inoculum containing both ErMF and asymbiotic saprobic fungi.
A representative of the new hyaloscyphoid clade repeatedly formed intracellular hyphal structures identical to those formed by typical ErMF and since it also supported the growth of the inoculated blueberry seedlings, there is a good chance that it represents a new ErMF, similarly to the recently described H. gryndleri (Vohník et al. 2022). However, this must be confirmed by more experimental studies, ideally employing other members of the clade and perhaps also isolates of the sexual saprobic H. daedaleae and H. fuckelii for comparison.
Conclusions
Rather than providing an exhaustive account of fungi inhabiting the roots of ericaceous plants growing on decomposing wood, this study offers a complex peek beyond the traditional scheme “ericoid mycorrhiza = acidic peaty substrates”, revealing once again how little we know about this important root-fungus association. Our observations do not support the view that typical EcM and saprobic basidiomycetes are mycorrhizal symbionts of ericaceous plants, but this issue is far from being solved and more experimental work is needed. Mountainous forested areas in Central Europe seem to be an unexpectedly rich reservoir of new root-symbiotic hyaloscyphoid fungi (Fehrer et al. 2019; Vohník et al. 2022, this study) and we encourage their research especially in the hitherto overlooked non-peat substrates. Such substrates are also found in the Southern Hemisphere (especially Australia and South Africa) where many ericaceous plants thrive in sandy soils where wood and sclerophyllous leaves represent major (often the only available) sources of nitrogen and phosphorus.
Data Availability
The sequences obtained in this study were deposited in GenBank at NCBI, herbarium specimens (as dried cultures) were deposited in the Herbarium of the Institute of Botany, Czech Academy of Sciences, Průhonice, Czechia (PRA), and living cultures were deposited in the collection of the Westerdijk Fungal Biodiversity Institute in Utrecht, the Netherlands (CBS).
References
Allen TR, Millar T, Berch SM, Berbee ML (2003) Culturing and direct DNA extraction find different fungi from the same ericoid mycorrhizal roots. New Phytol 160:255–272. https://doi.org/10.1046/j.1469-8137.2003.00885.x
Baba T, Hirose D (2021) Slow-growing fungi belonging to the unnamed lineage in Chaetothyriomycetidae form hyphal coils in vital ericaceous rhizodermal cells in vitro. Fungal Biol. https://doi.org/10.1016/j.funbio.2021.07.003
Baral H-O, De Sloover J, Huhtinen S, et al (2009) An emendation of the genus Hyaloscypha to include Fuscoscypha (Hyaloscyphaceae, Helotiales, Ascomycotina). Karstenia 49:1–17. https://doi.org/10.29203/ka.2009.430
Baucher M, Monties B, Van MM, Boerjan W (1998) Biosynthesis and genetic engineering of lignin. CRC Crit Rev Plant Sci 17:125–197. https://doi.org/10.1080/07352689891304203
Bending GD, Read DJ (1997) Lignin and soluble phenolic degradation by ectomycorrhizal and ericoid mycorrhizal fungi. Mycol Res 101:1348–1354. https://doi.org/10.1017/S0953756297004140
Boddy L, Watkinson SC (1995) Wood decomposition, higher fungi, and their role in nutrient redistribution. Can J Bot 73:1377–1383. https://doi.org/10.1139/b95-400
Bogale M, Orr M-J, O’Hara MJ, Untereiner WA (2010) Systematics of Catenulifera (anamorphic Hyaloscyphaceae) with an assessment of the phylogenetic position of Phialophora hyalina. Fungal Biol 114:396–409. https://doi.org/10.1016/j.funbio.2010.02.006
Bonfante-Fasolo P (1980) Occurrence of a basidiomycete in living cells of mycorrhizal hair roots of Calluna vulgaris. Trans Br Mycol Soc 75:320–325. https://doi.org/10.1016/S0007-1536(80)80097-0
Bougoure DS, Parkin PI, Cairney JWG et al (2007) Diversity of fungi in hair roots of Ericaceae varies along a vegetation gradient. Mol Ecol 16:4624–4636. https://doi.org/10.1111/j.1365-294X.2007.03540.x
Bruzone MC, Fontenla SB, Vohník M (2015) Is the prominent ericoid mycorrhizal fungus Rhizoscyphus ericae absent in the Southern Hemisphere’s Ericaceae? A case study on the diversity of root mycobionts in Gaultheria spp. from northwest Patagonia, Argentina. Mycorrhiza 25:25–40. https://doi.org/10.1007/s00572-014-0586-3
Cairney JWG, Burke RM (1998) Extracellular enzyme activities of the ericoid mycorrhizal endophyte Hymenoscyphus ericae (Read) Korf and Kernan: Their likely roles in decomposition of dead plant tissue in soil. Plant Soil 205:181–192. https://doi.org/10.1023/A:1004376731209
Chen C, Verkley GJM, Sun G et al (2016) Redefining common endophytes and plant pathogens in Neofabraea, Pezicula, and related genera. Fungal Biol 120:1291–1322. https://doi.org/10.1016/j.funbio.2015.09.013
Daghino S, Martino E, Voyron S, Perotto S (2022) Metabarcoding of fungal assemblages in Vaccinium myrtillus endosphere suggests colonization of above-ground organs by some ericoid mycorrhizal and DSE fungi. Sci Rep 12:11013. https://doi.org/10.1038/s41598-022-15154-1
Duclos JL, Pépin R, Bruchet G (1983) Étude morphologique, anatomique et ultrastructurale d’endomycorhizes synthétiques d’ Erica carnea. Can J Bot 61:466–475. https://doi.org/10.1139/b83-054
Fehrer J, Réblová M, Bambasová V, Vohník M (2019) The root-symbiotic Rhizoscyphus ericae aggregate and Hyaloscypha (Leotiomycetes) are congeneric: Phylogenetic and experimental evidence. Stud Mycol 92:195–225. https://doi.org/10.1016/j.simyco.2018.10.004
Fisher PJ, Anson AE, Petrini O (1984) Novel antibiotic activity of an endophytic Cryptosporiopsis sp. isolated from Vaccinium myrtillus. Trans Br Mycol Soc 83:145–148. https://doi.org/10.1016/S0007-1536(84)80254-5
Goodell B, Winandy JE, Morrell JJ (2020) Fungal degradation of wood: Emerging data, new insights and changing perceptions. Coatings 10:1210. https://doi.org/10.3390/coatings10121210
Gorzelak MA, Hambleton S, Massicotte HB (2012) Community structure of ericoid mycorrhizas and root-associated fungi of Vaccinium membranaceum across an elevation gradient in the Canadian Rocky Mountains. Fungal Ecol 5:36–45. https://doi.org/10.1016/j.funeco.2011.08.008
Gouy M, Guindon S, Gascuel O (2010) Seaview version 4: a multiplatform graphical user interface for sequence alignment and phylogenetic tree building. Mol Biol Evol 27:221–224. https://doi.org/10.1093/molbev/msp259
Grelet GA, Ba R, Goeke DF et al (2017) A plant growth-promoting symbiosis between Mycena galopus and Vaccinium corymbosum seedlings. Mycorrhiza 27:831–839. https://doi.org/10.1007/s00572-017-0797-5
Griffin A, Kernaghan G (2022) Ericoid mycorrhizal colonization and associated fungal communities along a wetland gradient in the Acadian forest of Eastern Canada. Fungal Ecol 56:101138. https://doi.org/10.1016/j.funeco.2021.101138
Grunewaldt-Stöcker G, von Alten H (2016) Is the root-colonizing endophyte Acremonium strictum an ericoid mycorrhizal fungus? Mycorrhiza 26:429–440. https://doi.org/10.1007/s00572-016-0682-7
Grünig CR, Queloz V, Sieber TN, Holdenrieder O (2008) Dark septate endophytes (DSE) of the Phialocephala fortinii s. l. – Acephala applanata species complex in tree roots: Classification, population biology, and ecology. Botany 86:1355–1369. https://doi.org/10.1139/B08-108
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Hambleton S, Currah RS (1997) Fungal endophytes from the roots of alpine and boreal Ericaceae. Can J Bot 75:1570–1581. https://doi.org/10.1139/b97-869
Hambleton S, Sigler L (2005) Meliniomyces, a new anamorph genus for root-associated fungi with phylogenetic affinities to Rhizoscyphus ericae (=Hymenoscyphus ericae), Leotiomycetes. Stud Mycol 53:1–27. https://doi.org/10.3114/sim.53.1.1
Han JG, Hosoya T, Sung GH, Shin HD (2014) Phylogenetic reassessment of Hyaloscyphaceae sensu lato (Helotiales, Leotiomycetes) based on multigene analyses. Fungal Biol 118:150–167. https://doi.org/10.1016/j.funbio.2013.11.004
Harder CB, Hesling E, Niskanen T et al (2021) Mycena species can be opportunist-generalist plant root invaders. bioRxiv. https://doi.org/10.1101/2021.03.23.436563
Huelsenbeck JP, Ronquist F (2001) MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics 17:754–755. https://doi.org/10.1093/bioinformatics/17.8.754
Janusz G, Pawlik A, Sulej J et al (2017) Lignin degradation: Microorganisms, enzymes involved, genomes analysis and evolution. FEMS Microbiol Rev 41:941–962. https://doi.org/10.1093/femsre/fux049
Katoh K, Standley DM (2013) MAFFT Multiple Sequence Alignment Software Version 7: Improvements in Performance and Usability. Mol Biol Evol 30:772–780. https://doi.org/10.1093/molbev/mst010
Kolařík M, Vohník M (2018) When the ribosomal DNA does not tell the truth: The case of the taxonomic position of Kurtia argillacea, an ericoid mycorrhizal fungus residing among Hymenochaetales. Fungal Biol 122:1–18. https://doi.org/10.1016/j.funbio.2017.09.006
Kosonen T, Huhtinen S, Hansen K (2021) Taxonomy and systematics of Hyaloscyphaceae and Arachnopezizaceae. Persoonia - Mol Phylogeny Evol Fungi 46:26–62. https://doi.org/10.3767/persoonia.2021.46.02
Kron KA, Judd WS, Stevens PF et al (2002) Phylogenetic classification of Ericaceae: Molecular and morphological evidence. Bot Rev 68:335–423. https://doi.org/10.1663/0006-8101(2002)068[0335:pcoema]2.0.co;2
Kron KA, Luteyn JL (2005) Origins and biogeographic patterns in Ericaceae: New insights from recent phylogenetic analyses. Biol Skr (Plant Divers Complex patterns local Reg Glob Dimens Proc an Int Symp held R Danish Acad Sci 1:479–500
Leake JR, Read DJ (1991) 20 Experiments with ericoid mycorrhiza. In: Norris JR, Read DJ, Varma AK (eds) Methods in Microbiology 23. Academic Press, London, pp 435–459
Lee Y-I, Yang C-K, Gebauer G (2015) The importance of associations with saprotrophic non-Rhizoctonia fungi among fully mycoheterotrophic orchids is currently under-estimated: novel evidence from sub-tropical Asia. Ann Bot 116:423–435. https://doi.org/10.1093/aob/mcv085
Leopold DR (2016) Ericoid fungal diversity: Challenges and opportunities for mycorrhizal research. Fungal Ecol 24:114–123. https://doi.org/10.1016/j.funeco.2016.07.004
Leopold DR, Peay KG, Vitousek PM, Fukami T (2021) Diversity of putative ericoid mycorrhizal fungi increases with soil age and progressive phosphorus limitation across a 4.1-million-year chronosequence. FEMS Microbiol Ecol 97:fiab016. https://doi.org/10.1093/femsec/fiab016
Lorberau KE, Botnen SS, Mundra S et al (2017) Does warming by open-top chambers induce change in the root-associated fungal community of the arctic dwarf shrub Cassiope tetragona (Ericaceae)? Mycorrhiza 27:513–524. https://doi.org/10.1007/s00572-017-0767-y
Lukešová T, Kohout P, Větrovský T, Vohník M (2015) The potential of dark septate endophytes to form root symbioses with ectomycorrhizal and ericoid mycorrhizal middle european forest plants. PLoS ONE. https://doi.org/10.1371/journal.pone.0124752
Martino E, Morin E, Grelet G et al (2018) Comparative genomics and transcriptomics depict ericoid mycorrhizal fungi as versatile saprotrophs and plant mutualists. New Phytol 217:1213–1229. https://doi.org/10.1111/nph.14974
Massicotte HB, Melville LH, Peterson RL (2005) Structural characteristics of root-fungal interactions for five ericaceous species in eastern Canada. Can J Bot 83:1057–1064. https://doi.org/10.1139/b05-046
Mayerhofer MS, Kernaghan G, Harper KA (2013) The effects of fungal root endophytes on plant growth: a meta-analysis. Mycorrhiza 23:119–128. https://doi.org/10.1007/s00572-012-0456-9
Midgley DJ, Jordan LA, Saleeba JA, McGee PA (2006) Utilisation of carbon substrates by orchid and ericoid mycorrhizal fungi from Australian dry sclerophyll forests. Mycorrhiza 16:175–182. https://doi.org/10.1007/s00572-005-0029-2
Midgley DJ, Rosewarne CP, Greenfield P et al (2016) Genomic insights into the carbohydrate catabolism of Cairneyella variabilis gen. nov. sp. nov., the first reports from a genome of an ericoid mycorrhizal fungus from the southern hemisphere. Mycorrhiza 26:345–352. https://doi.org/10.1007/s00572-016-0683-6
Midgley DJ, Sutcliffe B, Greenfield P, Tran-Dinh N (2018) Gamarada debralockiae gen. nov. sp. nov. – the genome of the most widespread Australian ericoid mycorrhizal fungus. Mycorrhiza 28:379–389. https://doi.org/10.1007/s00572-018-0835-y
Miller MA, Pfeiffer W, Schwartz T (2010) Creating the CIPRES Science Gateway for inference of large phylogenetic trees. In: 2010 Gateway Computing Environments Workshop (GCE). IEEE, pp 1–8
Molina R, Palmer JG (1982) Isolation, maintenance, and pure culture manipulation of ectomycorrhizal fungi. In: Schenck NC (ed) Methods and principles of mycorrhizal research. The American Phytopathological Society, St.Paul, Minnesota, pp 115–129
Newsham KK (2011) A meta-analysis of plant responses to dark septate root endophytes. New Phytol 190:783–793. https://doi.org/10.1111/j.1469-8137.2010.03611.x
Noble HM, Langley D, Sidebottom PJ et al (1991) An echinocandin from an endophytic Cryptosporiopsis sp. and Pezicula sp. in Pinus sylvestris and Fagus sylvatica. Mycol Res 95:1439–1440. https://doi.org/10.1016/S0953-7562(09)80401-2
Nylander JAA (2004) MrModeltest Version 2
Ogura-Tsujita Y, Gebauer G, Hashimoto T et al (2009) Evidence for novel and specialized mycorrhizal parasitism: the orchid Gastrodia confusa gains carbon from saprotrophic Mycena. Proc R Soc B Biol Sci 276:761–767. https://doi.org/10.1098/rspb.2008.1225
Pearson V, Read DJ (1975) The physiology of the mycorrhizal endophyte of Calluna vulgaris. Trans Br Mycol Soc 64:1–7. https://doi.org/10.1016/s0007-1536(75)80069-6
Perotto S, Daghino S, Martino E (2018) Ericoid mycorrhizal fungi and their genomes: another side to the mycorrhizal symbiosis? New Phytol 220:1141–1147. https://doi.org/10.1111/nph.15218
Perotto S, Girlanda M, Martino E (2002) Ericoid mycorrhizal fungi: Some new perspectives on old acquaintances. Plant Soil 244:41–53. https://doi.org/10.1023/A:1020289401610
Read DJ (1996) The structure and function of the ericoid mycorrhizal root. Ann Bot 77:365–374. https://doi.org/10.1006/anbo.1996.0044
Read DJ, Kerley S (1995) The status and function of ericoid mycorrhizal systems. Mycorrhiza. https://doi.org/10.1007/978-3-662-08897-5_22
Sayers EW, Cavanaugh M, Clark K et al (2019) GenBank. Nucleic Acids Res 47:D94–D99. https://doi.org/10.1093/nar/gky989
Schulz B, Boyle C (2006) What are endophytes? In: Schulz B, Boyle C, Sieber TN (eds) Microbial Root Endophytes. Springer, Berlin Heidelberg, Berlin, Heidelberg, pp 1–13
Schulz B, Sucker J, Aust HJ et al (1995) Biologically active secondary metabolites of endophytic Pezicula species. Mycol Res 99:1007–1015. https://doi.org/10.1016/S0953-7562(09)80766-1
Selosse MA, Setaro S, Glatard F et al (2007) Sebacinales are common mycorrhizal associates of Ericaceae. New Phytol 174:864–878. https://doi.org/10.1111/j.1469-8137.2007.02064.x
Setaro S, Weiß M, Oberwinkler F, Kottke I (2006) Sebacinales form ectendomycorrhizas with Cavendishia nobilis, a member of the Andean clade of Ericaceae, in the mountain rain forest of southern Ecuador. New Phytol 169:355–365. https://doi.org/10.1111/j.1469-8137.2005.01583.x
Sieber TN (2007) Endophytic fungi in forest trees: are they mutualists? Fungal Biol Rev 21:75–89. https://doi.org/10.1016/j.fbr.2007.05.004
Sigler L, Allan T, Lim SR et al (2005) Two new Cryptosporiopsis species from roots of ericaceous hosts in western North America. Stud Mycol 53:53–62. https://doi.org/10.3114/sim.53.1.53
Smith SE, Read D (2008) Ericoid mycorrhizas. In: Mycorrhizal Symbiosis. Elsevier, pp 389–418
Stamatakis A (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30:1312–1313. https://doi.org/10.1093/bioinformatics/btu033
Stenroos S, Laukka T, Huhtinen S, et al (2010) Multiple origins of symbioses between ascomycetes and bryophytes suggested by a five-gene phylogeny. Cladistics 26:281–300. https://doi.org/10.1111/j.1096-0031.2009.00284.x
Stillwell MA, Wood FA, Strunz GM (1969) A broad-spectrum antibiotic produced by a species of Cryptosporiopsis. Can J Microbiol 15:501–507. https://doi.org/10.1139/m69-087
Toju H, Tanabe AS, Ishii HS (2016) Ericaceous plant-fungus network in a harsh alpine-subalpine environment. Mol Ecol 25:3242–3257. https://doi.org/10.1111/mec.13680
Verkley GJM (1999) A monograph of the genus Pezicula and its anamorphs. Stud Mycol 5–180
Verkley GJM, Zijlstra JD, Summerbell RC, Berendse F (2003) Phylogeny and taxonomy of root-inhabiting Cryptosporiopsis species, and C. rhizophila sp. nov., a fungus inhabiting roots of several Ericaceae. Mycol Res 107:689–698. https://doi.org/10.1017/S0953756203007883
Villarreal-Ruiz L, Neri-Luna C, Anderson IC, Alexander IJ (2012) In vitro interactions between ectomycorrhizal fungi and ericaceous plants. Symbiosis 56:67–75. https://doi.org/10.1007/s13199-012-0161-7
Vohník M (2020) Ericoid mycorrhizal symbiosis: theoretical background and methods for its comprehensive investigation. Mycorrhiza 30:671–695. https://doi.org/10.1007/s00572-020-00989-1
Vohník M, Albrechtová J (2011) The co-occurrence and morphological continuum between ericoid mycorrhiza and dark septate endophytes in roots of six European Rhododendron species. Folia Geobot 46:373–386. https://doi.org/10.1007/s12224-011-9098-5
Vohník M, Albrechtová J, Vosátka M (2005) The inoculation with Oidiodendron maius and Phialocephala fortinii alters phosphorus and nitrogen uptake, foliar C: N ratio and root biomass distribution in Rhododendron cv. Azurro. Symbiosis 40:87–96
Vohník M, Figura T, Réblová M (2022) Hyaloscypha gabretae and Hyaloscypha gryndleri spp. nov. (Hyaloscyphaceae, Helotiales), two new mycobionts colonizing conifer, ericaceous and orchid roots. Mycorrhiza 32:105–122. https://doi.org/10.1007/s00572-021-01064-z
Vohník M, Lukančič S, Bahor E et al (2003) Inoculation of Rhododendron cv. Belle-Heller with two strains of Phialocephala fortinii in two different substrates. Folia Geobot 38:191–200. https://doi.org/10.1007/BF02803151
Vohník M, Mrnka L, Lukešová T et al (2013) The cultivable endophytic community of Norway spruce ectomycorrhizas from microhabitats lacking ericaceous hosts is dominated by ericoid mycorrhizal Meliniomyces variabilis. Fungal Ecol 6:281–292. https://doi.org/10.1016/j.funeco.2013.03.006
Vohník M, Pánek M, Fehrer J, Selosse M-A (2016) Experimental evidence of ericoid mycorrhizal potential within Serendipitaceae (Sebacinales). Mycorrhiza 26:831–846. https://doi.org/10.1007/s00572-016-0717-0
Vohník M, Sadowsky JJ, Kohout P et al (2012a) Novel root-fungus symbiosis in Ericaceae: Sheathed ericoid mycorrhiza formed by a hitherto undescribed basidiomycete with affinities to Trechisporales. PLoS ONE 7:e39524. https://doi.org/10.1371/journal.pone.0039524
Vohník M, Sadowsky JJ, Lukešová T et al (2012b) Inoculation with a ligninolytic basidiomycete, but not root symbiotic ascomycetes, positively affects growth of highbush blueberry (Ericaceae) grown in a pine litter substrate. Plant Soil 355:341–352. https://doi.org/10.1007/s11104-011-1106-2
Vrålstad T, Fossheim T, Schumacher T (2000) Piceirhiza bicolorata - The ectomycorrhizal expression of the Hymenoscyphus ericae aggregate? New Phytol 145:549–563. https://doi.org/10.1046/j.1469-8137.2000.00605.x
Vrålstad T, Myhre E, Schumacher T (2002) Molecular diversity and phylogenetic affinities of symbiotic rootassociated ascomycetes of the Helotiales in burnt and metal polluted habitats. New Phytol 155:131–148. https://doi.org/10.1046/j.1469-8137.2002.00444.x
Walker JF, Aldrich-Wolfe L, Riffel A et al (2011) Diverse helotiales associated with the roots of three species of arctic ericaceae provide no evidence for host specificity. New Phytol 191:515–527. https://doi.org/10.1111/j.1469-8137.2011.03703.x
Wang Z, Binder M, Hibbett DS (2005) Life history and systematics of the aquatic discomycete Mitrula (Helotiales, Ascomycota) based on cultural, morphological, and molecular studies. Am J Bot 92:1565–1574. https://doi.org/10.3732/ajb.92.9.1565
Welch D, Scott D, Doyle S (2000) Studies on the paradox of seedling rarity in Vaccinium myrtillus L. in NE Scotland. Bot J Scotl 52:17–30. https://doi.org/10.1080/03746600008684942
Yuan Z, Verkley GJM (2015) Pezicula neosporulosa sp. nov. (Helotiales, Ascomycota), an endophytic fungus associated with Abies spp. in China and Europe. Mycoscience 56:205–213. https://doi.org/10.1016/j.myc.2014.06.004
Zhang L, Chen J, Lv Y et al (2012) Mycena sp., a mycorrhizal fungus of the orchid Dendrobium officinale. Mycol Prog 11:395–401. https://doi.org/10.1007/s11557-011-0754-1
Zhang Z, Schwartz S, Wagner L, Miller W (2000) A Greedy Algorithm for Aligning DNA Sequences. J Comput Biol 7:203–214. https://doi.org/10.1089/10665270050081478
Zijlstra JD, Van’t Hof P, Baar J et al (2005) Diversity of symbiotic root endophytes of the Helotiales in ericaceous plants and the grass, Deschampsia flexuosa. Stud Mycol 53:147–162. https://doi.org/10.3114/sim.53.1.147
Acknowledgements
The authors wish to thank (chronologically) Erik Joner for stimulating our research of ericoid mycorrhizal fungi in Mid-Norway, Rolf Nestby for coordinating logistics during the sampling in Mid-Norway and providing access to the Bioforsk's laboratory in Kvithamar, Jesse J. Sadowsky for the help with the sample collection, microscopy and fungal isolation in Mid-Norway and Tereza Knoblochová (born Lukešová) and M. Clara Bruzone (¡muchas gracias!) for the help with the resynthesis experiments. Useful comments on the theoretical background of our study from two anonymous reviewers are greatly appreciated.
Funding
Open access publishing supported by the National Technical Library in Prague. This work has been supported by the Czech Science Foundation (GAČR 18-05886S) and the Institute of Botany, Czech Academy of Sciences (RVO 67985939).
Author information
Authors and Affiliations
Contributions
MV: concept of the study, obtaining resources, sampling, laboratory work, writing of the manuscript. MR: molecular analyses, writing of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Vohník, M., Réblová, M. Fungi in hair roots of Vaccinium spp. (Ericaceae) growing on decomposing wood: colonization patterns, identity, and in vitro symbiotic potential. Mycorrhiza 33, 69–86 (2023). https://doi.org/10.1007/s00572-023-01101-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00572-023-01101-z